 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
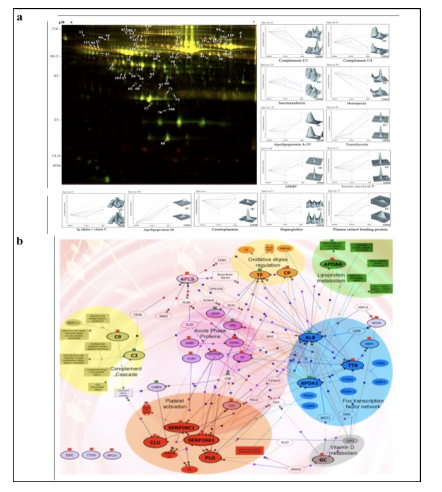
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)
 Shantikumar Nair, Ph.D.
Shantikumar Nair, Ph.D.
Professor & Director, Amrita Center for Nanosciences & Molecular Medicine, Amrita University, India
Spatially Distributed and Hierarchical Nanomaterials in Biotechnology
Although nano materials are well investigated in biotechnology in their zero-, one- and two-dimensional forms, three-dimensional nanomaterials are relatively less investigated for their biological applications. Three dimensional nano materials are much more complex with several structural and hierarchical variables controlling their mechanical, chemical and biological functionality. In this talk examples are given of some complex three dimensional systems including, scaffolds, aggregates, fabrics and membranes. Essentially three types of hierarchies are considered: one-dimensional hierarchy, two-dimensional hierarchy and three-dimensional hierarchy each giving rise to unique behaviors.
 Kal Ramnarayan, Ph.D.
Kal Ramnarayan, Ph.D.
Co-founder President & Chief Scientific Officer, Sapient Discovery, San Diego, CA, USA
A cost-effective approach to Protein Structure-guided Drug Discovery: Aided by Bioinformatics, Chemoinformatics and computational chemistry
With the mapping of the human genome completed almost a decade ago, efforts are still underway to understand the gene products (i.e., proteins) in the human biological and disease pathways. Deciphering such information is very important for the discovery and development of small molecule drugs as well as protein therapeutics for various human diseases for which no cure exists. As an example, with more than 500 members, the kinase family of protein targets continues to be an important and attractive class for drug discovery. While how many of the members in this family are actually druggable is still to be established, there are several ongoing efforts on this class of proteins across a broad spectrum of disease categories. Even though in general the protein structural topology might looks similar, there are issues with respect selectivity of identified small molecule inhibitors when, the lead molecule discovery is carried out at the ATP binding site. As an added complexity, allosteric modulators are needed for some of the members, but the actual site for such modulation on the protein target can not resolved with uncertainty. In this presentation we will describe a bioinformatics and computational based platform for small molecule discovery for protein targets that are involved in protein-protein interactions as well as targets like kinases and phosphatases. We will describe a computational approach in which we have used an informatics based platform with several hundred kinases to sort through in silico and identify inhibitors that are likely to be highly selective in the lead generation phase. We will discuss the implication of this approach on the drug discovery of the kinase and phosphatase classes in general and independent of the disease category.
 Manzoor K, Ph.D.
Manzoor K, Ph.D.
Professor, Centre for Nanoscience & Molecular Medicine, Amrita University
Targeting aberrant cancer kinome using rationally designed nano-polypharmaceutics
Manzoor Koyakutty, Archana Ratnakumary, Parwathy Chandran, Anusha Ashokan, and Shanti Nair
`War on Cancer’ was declared nearly 40 years ago. Since then, we made significant progress on fundamental understanding of cancer and developed novel therapeutics to deal with the most complex disease human race ever faced with. However, even today, cancer remains to be the unconquered `emperor of all maladies’. It is well accepted that meaningful progress in the fight against cancer is possible only with in-depth understanding on the molecular mechanisms that drives its swift and dynamic progression. During the last decade, emerging new technologies such as nanomedicine could offer refreshing life to the `war on cancer’ by way of providing novel methods for molecular diagnosis and therapy.
In the present talk, we discuss our approaches to target critically aberrant cancer kinases using rationally designed polymer-protein and protein-protein core-shell nanomedicines. We have used both genomic and proteomic approaches to identify many intimately cross-linked and complex aberrant protein kinases behind the drug resistance and uncontrolled proliferation of refractory leukemic cells derived from patients. Small molecule inhibitors targeted against oncogenic pathways in these cells were found ineffective due to the involvement of alternative survival pathways. This demands simultaneous inhibition more than one oncogenic kinases using poly-pharmaceutics approach. For this, we have rationally designed core-shell nanomedicines that can deliver several small molecules together for targeting multiple cancer signalling. We have also used combination of small molecules and siRNA for combined gene silencing together with protein kinase inhibition in refractory cancer cells. Optimized nanomedicines were successfully tested in patient samples and found enhanced cytotoxicity and molecular specificity in drug resistant cases.
Nano-polypharmaceutics represents a new generation of nanomedicines that can tackle multiple cancer mechanisms simultaneously. Considering the complexity of the disease, such therapeutic approaches are not simply an advantage, but indispensable.
Acknowledgements:
We thank Dept. of Biotechnology and Dept. Of Science and Technology,Govt. of India for the financial support through `Thematic unit of Excellence in Medical NanoBiotechnology’ and `Nanomedicine- RNAi programs’.
 Satheesh Babu T. G., Ph.D.
Satheesh Babu T. G., Ph.D.
Associate Professor, Department of Sciences, School of Engineering, Amrita University, Coimbatore, India
Nanomaterials for ‘enzyme-free’ biosensing
Enzyme based sensors have many draw backs such as poor storage stability, easily affected by the change in pH and temperature and involves complicated enzyme immobilization procedures. To address this limitation, an alternative approach without the use of enzyme, “non-enzymatic” has been tried recently. Choosing the right catalyst for direct electrochemical oxidation / reduction of a target molecule is the key step in the fabrication of non-enzymatic sensors.
Non-enzymatic sensors for glucose, creatinine, vitamins and cholesterol are fabricated using different nanomaterials, such as nanotubes, nanowires and nanoparticles of copper oxide, titanium dioxide, tantalum oxide, platinum, gold and graphenes. These sensors selectively catalyse the targeted analyte with very high sensitivity. These nanomaterials based sensors combat the drawbacks of enzymatic sensors.
![Delegate Talk: Pharmacophore modeling, atom-based 3D-QSAR and molecular docking studies on Pyrimido[5,4-e][1,2,4]triazine derivatives as PLK 1 inhibitors @ Sathyam Hall | Vallikavu | Kerala | India](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/bicb.jpg)
Rajasekhar Chekkara, Venkata Reddy Gorla and Sobha Rani Tenkayala
Pharmacophore modeling, atom-based 3D-QSAR and molecular docking studies on Pyrimido[5,4-e][1,2,4]triazine derivatives as PLK 1 inhibitors
Polo-like kinase 1 (PLK1) is a significant enzyme with diverse biological actions in cell cycle progression, specifically mitosis. Suppression of PLK1 activity by small molecule inhibitors has been shown to inhibit cancer, being BI 2536 one of the most potent active inhibitor of PLK1 mechanism. Pharmacophore modeling, atom-based 3D-QSAR and molecular docking studies were carried out for a set of 54 compounds belonging to Pyrimido[5,4-e][1,2,4]triazine derivatives as PLK1 inhibitors. A six-point pharmacophoremodel AAADDR, with three hydrogen bond acceptors (A), two hydrogen bond donors (D) and one aromatic ring (R) was developed by Phase module of Schrdinger suite Maestro 9. The generated pharmacophore model was used to derive a predictive atom-based 3D quantitative structure-activity relationship analysis (3D-QSAR) model for the training set (r2 = 0.88, SD = 0.21, F = 57.7, N = 44) and for test set (Q2 = 0.51, RMSE = 0.41, PearsonR = 0.79, N = 10). The original set of compounds were docked into the binding site of PLK1 using Glide and the active residues of the binding site were analyzed. The most active compound H18 interacted with active residues Leu 59, Cys133 (glide score = −10.07) and in comparison of BI 2536, which interacted with active residues Leu 59, Cys133 (glide score = −10.02). The 3D-QSAR model suggests that hydrophobic and electron-withdrawing groups are essential for PLK1 inhibitory activity. The docking results describes the hydrogen bond interactions with active residues of these compounds. These results which may support in the design and development of novel PLK1 inhibitors.
 Sudarslal S, Ph.D.
Sudarslal S, Ph.D.
Associate Professor, School of Biotechnology, Amrita University
Electrospray ionization ion trap mass spectrometry for cyclic peptide characterization
There has been considerable interest in the isolation and structural characterization of bioactive peptides produced by bacteria and fungi. Most of the peptides are cyclic depsipeptides characterized by the presence of lactone linkages and β-hydroxy fatty acids. Occurrence of microheterogeneity is another remarkable property of these peptides. Even if tandem mass spectrometers are good analytical tools to structurally characterize peptides and proteins, sequence analysis of cyclic peptides is often ambiguous due to the random ring opening of the peptides and subsequent generation of a set of linear precursor ions with the same m/z. Here we report combined use of chemical derivatization and multistage fragmentation capability of ion trap mass spectrometers to determine primary sequences of a series of closely related cyclic peptides.

Ravindra Gudihal, Suresh Babu C V
Bioanalytical Characterization of Therapeutic Proteins
The characterization of therapeutic proteins such as monoclonal antibody (mAb) during different stages of manufacturing is crucial for timely and successful product release. Regulatory agencies require a variety of analytical technologies for comprehensive and efficient protein analysis. Electrophoresis-based techniques and liquid chromatography (LC) either standalone or coupled to mass spectrometry (MS) are at the forefront for the in-depth analysis of protein purity, isoforms, stability, aggregation, posttranslational modifications, PEGylation, etc. In this presentation, a combination of various chromatographic and electrophoretic techniques such as liquid-phase isoelectric focusing, microfluidic and capillary-based electrophoresis (CE), liquid chromatography (LC) and combinations of those with mass spectrometry techniques will be discussed. We present a workflow based approach to the analysis of therapeutic proteins. In successive steps critical parameters like purity, accurate mass, aggregation, peptide sequence, glycopeptide and glycan analysis are analyzed. In brief, the workflow involved proteolytic digestion of therapeutic protein for peptide mapping, N-Glycanase and chemical labeling reaction for glycan analysis, liquid-phase isoelectric focusing for enrichment of charge variants followed by a very detailed analysis using state of the art methods such as CE-MS and LC-MS. For the analysis of glycans, we use combinations of CE-MS and LC-MS to highlight the sweet spots of these techniques. CE-MS is found to be more useful in analysis of highly sialylated glycans (charged glycans) while nano LC-MS seems to be better adapted for analysis of neutral glycans. These two techniques can be used to get complementary data to profile all the glycans present in a given protein. In addition, microfluidic electrophoresis was used as a QC tool in initial screening for product purity, analysis of papain digestion fragments of mAb, protein PEGylation products, etc. The described workflow involves multiple platforms, provides an end to end solution for comprehensive protein characterization and aims at reducing the total product development time.

Tejaswini Subbannayya, Nandini A. Sahasrabuddhe, Arivusudar Marimuthu, Santosh Renuse, Gajanan Sathe, Srinivas M. Srikanth, Mustafa A. Barbhuiya, Bipin Nair, Juan Carlos Roa, Rafael Guerrero-Preston, H. C. Harsha, David Sidransky, Akhilesh Pandey, T. S. Keshava Prasad and Aditi Chatterjee
Proteomic profiling of gallbladder cancer secretome – a source for circulatory biomarker discovery
Gallbladder cancer (GBC) is the fifth most common cancer of the gastrointestinal tract and one of the common malignancies that occur in the biliary tract (Misra et al. 2006; Lazcano-Ponce et al. 2001). It has a poor prognosis with survival of less than 5 years in 90% of the cases (Misra et al. 2003). The etiology is ill-defined. Several risk factors have been reported including cholelithiasis, obesity, female gender and exposure to carcinogens (Eslick 2010; Kumar et al. 2006). Poor prognosis in GBC is mainly due to late presentation of the disease and lack of reliable biomarkers for early diagnosis. This emphasizes the need to identify and characterize cancer biomarkers to aid in the diagnosis and prognosis of GBC. Secreted proteins are an important class of molecules which can be detected in body fluids and has been targeted for biomarker discovery. There are challenges faced in the proteomic interrogation of body fluids especially plasma such as low abundance of tumor secreted proteins, high complexity and high abundance of other proteins that are not released by the tumor cells (Tonack et al. 2009). Profiling of conditioned media from the cancer cell lines can be used as an alternate means to identify secreted proteins from tumor cells (Kashyap et al. 2010; Marimuthu et al. 2012). We analyzed the invasive property of 7 GBC cell lines (SNU-308, G-415, GB-d1, TGBC2TKB, TGBC24TKB, OCUG-1 and NOZ). Four cell lines were selected for analysis of the cancer secretome based on the invasive property of the cells. We employed isobaric tags for relative and absolute quantitation (iTRAQ) labeling technology coupled with high resolution mass spectrometry to identify and characterize secretome from the panel of 4GBC cancer cells mentioned above. In total, we have identified around 2,000 proteins of which 175 were secreted at differential abundance across all the four cell lines. This secretome analysis will act as a reservoir of candidate biomarkers. Currently, we are investigating and validating these candidate markers from GBC cell secretome. Through this study, we have shown mass spectrometry-based quantitative proteomic analysis as a robust approach to investigate secreted proteins in cancer cells.







