Dr. Bipin Nair,
Dean-Biotechnology, Amrita University
 Sudhir Sahasrabudhe, Ph.D.
Sudhir Sahasrabudhe, Ph.D.
CEO, Rines Therapeutics
Research Professor of Medicinal Chemistry, University of Utah, USA
NOVEL DRUG TARGETS AND NOVEL DRUGS USING PROTEOMICS TECHNOLOGY
Translational medicine is the conversion of scientific discovery into improvement of human health. The key to any translational medicine effort is to make sure that the novel inter-disciplinary research advances in the laboratory find their way to the clinic to better serve patients with unmet medical need. This talk will feature 2 such efforts:
Example 1:
We established a chemi-proteomics platform to capture and identify proteins that bind to small molecule hits from a cell-based genetic screen. This screen was geared to identify molecules that are selectively lethal to cells with aberrantly active RAS-signaling pathway (Nature: 447, 865, 2007; Nature Methods:7, 801, 2010). This effort helped identification of a novel cancer target and paved the way for creation of a novel small molecule currently in clinical Proof-of-concept (Phase 2a) studies in cancer patients
Example 2:
Huntington’s disease (HD) is a fatal neurodegenerative condition caused by expansion of the polyglutamine tract in the huntingtin (Htt) protein. Neuronal toxicity in HD is thought to be a consequence of protein interactions involving mutant Htt. We therefore hypothesized that genetic modifiers of HD neurodegeneration should be enriched among Htt protein interactors (PLOS Genetics: 3(5), 2007). To test this idea, we identified a comprehensive set of Htt interactors using two complementary approaches: high-throughput yeast two-hybrid screening and affinity pull down followed by mass spectrometry. This effort led to the identification of 234 high-confidence Htt-associated proteins, 104 of which were found with the yeast method and 130 with the pull downs. We then tested an arbitrary set of 60 genes encoding interacting proteins for their ability to behave as genetic modifiers of neurodegeneration in a Drosophila model of HD. This presentation will outline the identification and validation of novel drug targets for HD.
 Hideaki Nagase, Ph.D.
Hideaki Nagase, Ph.D.
Kennedy Institute of Rheumatology-Centre for Degenerative Diseases, University of Oxford, UK
Osteoarthritis: diagnosis, treatment and challenges
Hideaki Nagase1, Ngee Han Lim1, George Bou-Gharios1, Ernst Meinjohanns2 and Morten Meldal3
- Kennedy Institute of Rheumatology, Nuffield Department of Orthopaedics, Rheumatology and Musculoskeletal Sciences, University of Oxford, London, W6 8LH UK
- Carlsberg Laboratory, Copenhagen, Denmark,
- Nano-Science Center, Department of Chemistry, University of Copenhagen, Denmark
Osteoarthritis (OA) is the most prevalent age-related degenerative joint disease. With the expanding ageing population, it imposes a major socio-economic burden on society. A key feature of OA is a gradual loss of articular cartilage and deformation of bone, resulting in the impairment of joint function. Currently, there is no effective disease-modifying treatment except joint replacement surgery. There are many possible causes of cartilage loss (e.g. mechanical load, injury, reactive oxygen species, aging, etc.) and etiological factors (obesity, genetics), but the degradation of cartilage is primarily caused by elevated levels of active metalloproteinases. It is therefore attractive to consider proteinase inhibitors as potential therapeutics. However, there are several hurdles to overcome, namely early diagnosis and continuous monitoring of the efficacy of inhibitor therapeutics. We are therefore aiming at developing non-invasive probes to detect cartilage degrading metalloproteinase activities.
We have designed in vivo imaging probes to detect MMP-13 (collagenase 3) activity that participates in OA by degrade cartilage collagen II and MMP-12 (macrophage elastase) activity involved in inflammatory arthritis. These activity-based probes consist of a peptide that is selectively cleaved by the target proteinase, a near-infrared fluorophore and a quencher. The probe’s signal multiplies upon proteolysis. They were first used to follow the respective enzyme activity in vivo in the mouse model of collagen-induced arthritis and we found MMP-12 activity probe (MMP12AP) activation peaked at 5 days after onset of the disease, whereas MMP13AP activation was observed at 10-15 days. The in vivo activation of these probes was inhibited by specific low molecule inhibitors. We proceeded to test both probes in the mouse model of OA induced by the surgical destabilization of medial meniscus of the knee joints. In this model, degradation of knee cartilage is first detected histologically 6 weeks after surgery with significant erosion detectable at 8 weeks. Little activation of MMP12AP was detected, which was expected, as macrophage migration is not obvious in OA. MMP13AP, on the other hand, was significantly activated in the operated knee at 6 weeks compared with the non-operated contralateral knee, but there were no significant differences between the operated and sham-operated knees. At 8 weeks, however, the signals in the operated knees were significantly higher than both the contralateral and sham-operated controls. Activation of aggrecanases and MMP-13 are observed before structural changes of cartilage. We are therefore currently improving the MMP-13 probe for earlier detection by attaching it to polymers that are retained in cartilage.
 K. Satyamoorthy, Ph.D.
K. Satyamoorthy, Ph.D.
Director, Life Sciences Centre, Manipal University, India
Epigenetic Changes due to DNA Methylation in Human Epithelial Tumors
Extensive global hypomethylation in the genome and hypermthylation of selective tumor specific suppressor genes appears to be a hallmark of human cancers. Data suggests that hypermethylation of promoter region in genes is more closely related to subsequent gene expression; contrary to gene-body DNA methylation. The intricate balance between these two may contribute to the progressive process of development, differentiation and carcinogenesis. Epigenetic changes encompass, apart from DNA methylation, chromatin modifications through post-translational changes in histones and control by miRNAs. At the genome level, effects from these are compounded by copy number variations (CNVs) which may ultimately influence protein functions. From clinical perspective, changes in DNA methylation occur very early which are reversible and are influenced by environmental factors. Therefore, these can be potential resource for identifying therapeutic targets as well as biomarkers for early screening of cancer. Our current efforts in profiling genome wide DNA methylation changes in oral, cervical and breast cancers through DNA methylation microarray analysis has revealed number of alterations critical for survival, progression and metastatic behavior of tumors. Bioinformatics and functional analysis revealed several key regulatory molecules controlled by DNA methylation and suggests that DNA methylation changes in several CpG islands appear to co-segregate in the regions of miRNAs as well as in the CNVs. We have validated the signatures for methylation of CpG islands through bisufite sequencing for essential genes in clinical samples and have undertaken transcriptional and functional analysis in tumor cell lines. These results will be presented.
 Gillian Murphy, Ph.D.
Gillian Murphy, Ph.D.
Professor, Department of Oncology, University of Cambridge, UK
A novel strategy for targeting metalloproteinases in cancer
Epithelial tumours evolve in a multi-step manner, involving both inflammatory and mesenchymal cells. Although intrinsic factors drive malignant progression, the influence of the micro-environment of neoplastic cells is a major feature of tumorigenesis. Extracellular proteinases, notably the metalloproteinases, are key players in the regulation of this cellular environment, acting as major effectors of both cell-cell and cell-extracellular matrix (ECM) interactions. They are involved in modifying ECM integrity, growth factor availability and the function of cell surface signalling systems, with consequent effects on cellular differentiation, proliferation and apoptosis.This has made metalloproteinases important targets for therapeutic interventions in cancer and small molecule inhibitors focussed on chelation of the active site zinc and binding within the immediate active site pocket were developed. These were not successful in early clinical trials due to the relative lack of specificity and precise knowledge of the target proteinase(s) in specific cancers. We can now appreciate that it is essential that we understand the relative roles of the different enzymes (of which there are over 60) in terms of their pro and anti tumour activity and their precise sites of expression The next generations of metalloproteinase inhibitors need the added specificity that might be gained from an understanding of the structure of individual active sites and the role of extra catalytic domains in substrate binding and other aspects of their biology. We have prepared scFv antibodies to the extra catalytic domains of two membrane metalloproteinases, MMP-14 and ADAM17, that play key roles in the tumour microenvironment. Our rationale and experiences with these agents will be presented in more detail.
 Terry Hermiston, Ph.D.
Terry Hermiston, Ph.D.
Vice President, US Biologics Research Site Head, US Innovation Center Bayer Healthcare, USA
ColoAd1 – An oncolytic adenovirus derived by directed evolution
Attempts at developing oncolytic viruses have been primarily based on rational design. However, this approach has been met with limited success. An alternative approach employs directed evolution as a means of producing highly selective and potent anticancer viruses. In this method, viruses are grown under conditions that enrich and maximize viral diversity and then passaged under conditions meant to mimic those encountered in the human cancer microenvironment. Using the “Directed Evolution” methodology, we have generated ColoAd1, a novel chimeric oncolytic adenovirus. In vitro, this virus demonstrated a >2 log increase in both potency and selectivity when compared to ONYX-015 on colon cancer cells. These results were further supported by in vivo and ex vivo studies. Importantly, these results have validated this methodology as a new general approach for deriving clinically-relevant, highly potent anti-cancer virotherapies. This virus is currently in clinical trials as a novel treatment for cancer.
 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
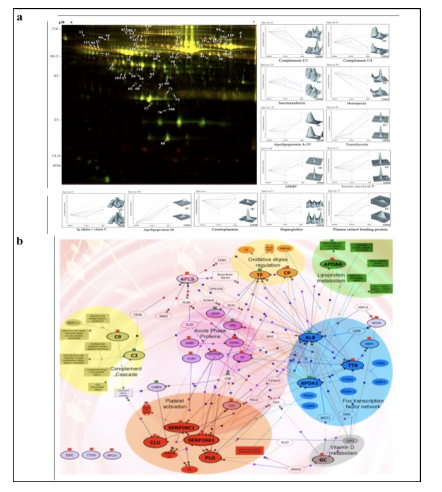
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)

Anna Travesa, Curt Wittenerg, Dwight Kuo, Robert A.M. de Bruin, Trey Ideker and Curt Wittenberg
Genomic and Genetic Analyses of the Transcriptional Regulation of G1/S Genes by Genotoxic Stress
Anna Travesa1,4, Dwight Kuo2, Robert A.M. de Bruin3, Trey Ideker2, and Curt Wittenberg1
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037
- Departments of Bioengineering and Medicine,University of California, San Diego, La Jolla, CA 92093
- MRC Laboratory for Molecular Cell Biology, University College London, London, WC1E 6BT United Kingdom
- Current address: Department of Cell and Developmental Biology, University of California, San Diego, La Jolla, CA 92093
The G1/S-phase transition is critical for cell cycle progression. In the budding yeast Saccharomyces cerevisiae, this transition is driven by a wave of gene expression that is under the control of the cell cycle-regulated transcription factors SBF and MBF. SBF mainly targets genes involved in timing, morphogenesis and spindle pole body duplication, whereas MBF regulates many genes involved in DNA replication and repair. During S phase, the DNA replication checkpoint detects DNA replication stress and generates a global response that includes the transcriptional induction of genes involved in DNA replication and repair. We find that MBF-dependent transcription is induced in response to DNA damage and replication stress. This induction takes place via direct phosphorylation of the transcriptional corepressor Nrm1 by the checkpoint kinase Rad53, which renders it unable to bind to promoters and repress transcription as cells progress into S phase. Using genome-wide RNA microarrays, we have shown that approximately a third of the G1/S specific genes are induced in response to DNA damage and replication stress, and that this induction is largely dependent upon Rad53. Moreover, almost 80 % of those genes are regulated by Nrm1, highlighting the general relevance of the Rad53-dependent inactivation of Nrm1 by the checkpoint. This checkpoint regulation of G1/S gene expression is conserved in the distantly related fungi, Schizosaccharomyces pombe, and also in eukaryotic cells. By inducing many genes involved in DNA replication and repair, this newly characterized pathway enhances genomic stability in the face of a broad range of genotoxic stresses.

Ravi Gutti, Rambabu Undi, Ravinder Kandi and Itishri Sahu
Hematologic Oncology, Stem Cells and Blood Disorders Laboratory, School of Life Sciences, Department of Biochemistry, University of Hyderabad, Gachibowli, Hyderabad 500 046 (AP), India.
e-mail: guttiravi@gmail.com
Megakaryocytopoiesis is governed by a complex network of haematopoietic growth factors that regulate the different stages of the process, in which haematopoietic stem cells undergo megakaryocytic lineage commitment, proliferation, maturation, and functional activation to produce platelets. Hyperproliferative progenitors and small, low-ploidyMKs in neonates may be due to the developmental differences in megakaryocytes (MKs) between neonates and adults. The biological relevance of these differences is highlighted by the fact that a number of disorders of megakaryocytopoiesis are restricted to the fetal and neonatal stages of development. The regulatory mechanisms underlying these developmental differences are unknown and the small non-codingmicroRNAs (miRNAs) shown to play a critical role in the regulation of MK development. We hypothesized that miRNAs would be differentially expressed in neonatal and adult MKs, and that these differences would contribute to their biological differences. To test this, we cultured human cord blood (CB) and peripheral blood (PB) CD34+ cells in serum free media with thrombopoietin. After 14 days of culture, > 90% of the cells were MKs (CD41+). miRNA and protein was isolated and expression levels of 88 miRNAs known to be involved in human stem cell differentiation and development were measured using a quantitative PCR-based array kit. Web-based computational approaches (TargetScan, PicTar and MiRanda) were used for putative target prediction and the protein levels were detected using western blot analysis. All samples (n = 4 per group) expressed detectable amounts of all 88 screened miRNAs. TenmiRNAs were expressed at significantly higher levels in CB compared toPBMKs (2 to 21 fold, p < 0.05. We then looked for the target for thisupregulatedmiRNAs in CB MKs. RUNX-1 was a predicted target of five of theupregulatedmiRNAs in CBMKs (miR-9, miR-129-5p, miR-192, miR-215, and miR-370). RUNX1 is an eukaryotic gene and the protein encoded by this gene is a transcription factor, also called AML-1 which is critical for MK maturation. In humans, loss of function mutations in RUNX-1 cause familial platelet disorder with propensity to develop acute myeloid leukemia, an autosomal dominant disorder characterized by quantitative and qualitative platelet defects and a predisposition to develop AML. In addition, RUNX 1 protein levels were approximately 3-fold higher inPB compared to CB MKs. Our results indicate that there are significant differences in the RUNX-1 expression and decreased RUNX-1 levels are consistent with the phenotype of increased proliferation and decreased size andploidy that characterizes neonatal megakaryopoiesis. Low RUNX-1 protein levels in neonatal compared to adult MKs, suggests that its translation is inhibited by those overexpressed miRNAs. We found activation of cyclin D3 transcription and repression of the p21 promoter likely contribute to stimulation of proliferation by Runx1. We believe that Wnt ligands could also stimulate cyclin D and c-Myc expression to favor cell proliferation by stimulating G1 entry from G0 and G1 to S cell cycle progression (Friedman 2009). Epigenetic/transcriptional control, RNA processing, and regulation of protein stability also likely to influence Runx1 activities in developmentalmegakaryocytopoiesis. Further studies to determine whether this contributes to the neonatal MK phenotype are in progress.

Binu K Aa, Jem Prabhakarb, Thara Sc and Lakshmi Sd,∗
aDepartment of Clinical Diagnostics Services and Translational Research, Malabar Cancer Centre, Thalassery, Kerala, India.
bDivision of Surgical Oncology, Division of Pathology
dDivision of Cancer Research, Regional Cancer Centre, Thiruvananthapuram, Kerala, India.
Introduction
AIB1, a member of the nuclear co activators, promotes the transcriptional activity of multiple nuclear receptors such as the ER and other transcription factors. Chemokines produced by stromal cells have potential to influence ERα-positive breast cancer progression to metastasis. CXCR4 is the physiological receptor for SDF1, together shown to stimulate the chemotactic and invasive behavior of breast cancer cells to serve as a homing mechanism to sites of metastasis. We propose that over expression of AIB1 in breast cancer cells leads to increased SDF1 and CXCR4 expression, which induces invasion and metastasis of cancer cells.
Materials and Methods
Breast tumor and normal breast tissues from patients in Regional Cancer Centre, Thiruvananthapuram were used for study. The modulatory effect of AIB1 was studied in MCF-7 cells with AIB1 siRNA transfection along with treatment of 17β-Estradiol (E2), 4-hydroxytamoxifen (4OHT), combinations of E2 and 4OHT. The gene expression pattern and protein localization were assessed by RT-PCR and immunofluorescence microscopy respectively. The metastatic and invasive properties were assessed by wound healing assay. Quantitative colocalization analyses were done to assess the association of proteins using Pearson’s correlation coefficient.
Result and Conclusion
The mRNA and protein level expression of AIB1, CXCR4 and SDF1 were higher in tumor samples than in normal samples. AIB1 was localized to the nuclei whereas CXCR4 and SDF1 immunoreactivity were observed in the cytoplasm and to a lesser extent in the nuclei of tumor epithelial cells. In tumor samples the gene level expressions of AIB1 showed significant positive correlations with SDF1(r = 0.213, p = 0.018). CXCR4 showed significant positive correlation with SDF1 in gene (r = 0.498, p = 0.000) and protein levels(r = 0.375, p = 0.002). Quantitative colocalization analyses showed a marked reduction in expression of CXCR4 and SDF1 in siAIB1MCF-7 cells than MCF-7 cells with different treatment groups. Wound healing assay shows reduced wound healing in siAIB1 treated MCF-7 cells.
In recent years, targeting specific cancer pathways and key molecules to arrest tumor growth and achieve tumor eradication have proven a challenge; due to acquired resistance and homing of cancer cells to various metastatic sites. The present study revealed that silencing AIB1 can prevent the over expression of SDF1 and CXCR4. Co activator levels determine the basal and estrogen-inducible expression of SDF1, a secreted protein that controls breast cancer cell proliferation and invasion through autocrine and paracrine mechanisms (Hall et al. 2003). The effects of CXCR4 overexpression has been correlated with SDF1 mediated activation of downstream signaling via ERK1/2 and p38 MAPK and with an enhancement of ER-mediated gene expression (Rhodes et al. 2011). It is possible that over expression of AIB1 as a stimulant involved in the expression of CXCR4 might up-regulate the expression of prometastatic and angiogenic genes. Thus based on these observations it can be concluded that SDF1/CXCR4 overexpression, with significant association with AIB1 expression, itself contribute to the development of mammary cancer and metastatic progression.
 Kartik Iyer
Kartik Iyer
Assistant Professor, School of Biotechnology, Amrita University
 Suryaprakash Sambhara, DVM, Ph.D
Suryaprakash Sambhara, DVM, Ph.D
Chief, Immunology Section, Influenza Division, CDC, Atlanta, USA
Making sense of pathogen sensors of Innate Immunity: Utility of their ligands as antiviral agents and adjuvants for vaccines.
Currently used antiviral agents act by inhibiting viral entry, replication, or release of viral progeny. However, recent emergence of drug-resistant viruses has become a major public health concern as it is limiting our ability to prevent and treat viral diseases. Furthermore, very few antiviral agents with novel modes of action are currently in development. It is well established that the innate immune system is the first line of defense against invading pathogens. The recognition of diverse pathogen-associated molecular patterns (PAMPs) is accomplished by several classes of pattern recognition receptors (PRRs) and the ligand/receptor interactions trigger an effective innate antiviral response. In the past several years, remarkable progress has been made towards understanding both the structural and functional nature of PAMPs and PRRs. As a result of their indispensable role in virus infection, these ligands have become potential pharmacological agents against viral infections. Since their pathways of action are evolutionarily conserved, the likelihood of viruses developing resistance to PRR activation is diminished. I will discuss the recent developments investigating the potential utility of the ligands of innate immune receptors as antiviral agents and molecular adjuvants for vaccines.
 D. Narasimha Rao, Ph.D.
D. Narasimha Rao, Ph.D.
Professor, Dept of Biochemistry, Indian Institute of Science, Bangalore, India
Genomics of Restriction-Modification Systems
Restriction endonucleases occur ubiquitously among procaryotic organisms. Up to 1% of the genome of procaryotic organisms is taken up by the genes for these enzymes. Their principal biological function is the protection of the host genome against foreign DNA, in particular bacteriophage DNA. Restriction-modification (R-M) systems are composed of pairs of opposing enzyme activities: an endonuclease and a DNA methyltransferase (MTase). The endonucleases recognise specific sequences and catalyse cleavage of double-stranded DNA. The modification MTases catalyse the addition of a methyl group to one nucleotide in each strand of the recognition sequence using S-adenosyl-L-methionine (AdoMet) as the methyl group donor. Based on their molecular structure, sequence recognition, cleavage position and cofactor requirements, R-M systems are generally classified into three groups. In general R-M systems restrict unmodified DNA, but there are other systems that specifically recognise and cut modified DNA. More than 3500 restriction enzymes have been discovered so far. With the identification and sequencing of a number of R-M systems from bacterial genomes, an increasing number of these have been found that do not seem to fit into the conventional classification.
It is well documented that restriction enzyme genes always lie close to their cognate methyltransferase genes. Analysis of the bacterial and archaeal genome sequences shows that MTase genes are more common than one would have expected on the basis of previous biochemical screening. Frequently, they clearly form part of a R-M system, because the adjacent open reading frames (ORFs) show similarity to known restriction enzyme genes. Very often, though, the adjacent ORFs have no homologs in the GenBank and become candidates either for restriction enzymes with novel specificities or for new examples of previously uncloned specificities. Sequence-dependent modification and restriction forms the foundation of defense against foreign DNAs and thus RM systems may serve as a tool of defense for bacterial cells. RM systems however, sometimes behave as discrete units of life, and any threat to their maintenance, such as a challenge by a competing genetic element can lead to cell death through restriction breakage in the genome, thus providing these systems with a competitive advantage. The RM systems can behave as mobile-genetic elements and have undergone extensive horizontal transfer between genomes causing genome rearrangements. The capacity of RM systems to act as selfish, mobile genetic elements may underlie the structure and function of RM enzymes.
The similarities and differences in the different mechanisms used by restriction enzymes will be discussed. Although it is not clear whether the majority of R-M systems are required for the maintenance of the integrity of the genome or whether they are spreading as selfish genetic elements, they are key players in the “genomic metabolism” of procaryotic organisms. As such they deserve the attention of biologists in general. Finally, restriction enzymes are the work horses of molecular biology. Understanding their enzymology will be advantageous to those who use these enzymes, and essential for those who are devoted to the ambitious goal of changing the properties of these enzymes, and thereby make them even more useful.
 Michelle Hermiston, MD, Ph.D.
Michelle Hermiston, MD, Ph.D.
Assistant Professor, Department of Pediatrics University of California San Francisco, USA
Interrogating Signaling Networks at the Single Cell Level In Primary Human Patient Samples
Multiparameter phosphoflow cytometry is a highly sensitive proteomic approach that enables monitoring of biochemical perturbations at the single cell level. By combining antisera to cell surface markers and key intracellular proteins, perturbations in signaling networks, cell survival and apoptosis mediators, cell cycle regulators, and/or modulators of other cellular processes can be analyzed in a highly reproducible and sensitive manner in the basal state and in response to stimulation or drug treatment. Advantages of this approach include the ability to identify the biochemical consequences of genetic and/or epigenetic changes in small numbers of cells, to map potential interplay between various signaling networks simultaneously in a single cell, and to interrogate potential mechanisms of drug resistance or response in a primary patient sample. Application of this technology to patients with acute lymphoblastic leukemia or the autoimmune disease systemic lupus erythematosus (SLE) will be discussed.
 Shigeki Miyamoto, Ph.D.
Shigeki Miyamoto, Ph.D.
Professor, McArdle Laboratory for Cancer Research – UW Carbone Cancer Center
Department of Oncology, School of Medicine and Public Health
University of Wisconsin-Madison
“Inside-out” NF-κB signaling in cancer and other pathologies
The NF-κB/Rel family of transcription factors contributes to critical cellular processes, including immune, inflammatory and cell survival responses. As such, NF-κB is implicated in immunity-related diseases, as well as multiple types of human malignancies. Indeed, genetic alterations in the NF-κB signaling pathway are frequently observed in multiple human malignancies. NF-κB is normally kept inactive in the cytoplasm by inhibitor proteins. Extracellular ligands can induce the release of NF-κB from the inhibitors to allow its migration into the nucleus to regulate a variety of target genes. NF-κB activation is also induced in response to multiple stress conditions, including those induced by DNA-damaging anticancer agents. Although precise mechanisms are still unclear, research from our group has revealed a unique nuclear-to-cytoplasmic signaling pathway. In collaboration with bioengineers, clinicians and pharmaceutical industry, our lab has developed new methods to analyze primary cancer patient samples and identified several compounds with different mechanisms that mitigate this cell survival pathway. Further contributions from other labs have also revealed additional mechanisms and molecular players in this “inside-out” signaling pathway and expanded its role in other physiological and pathological processes, including B cell development, premature aging and therapy resistance of certain cancers. Our own new findings, along with these recent developments in the field, will be highlighted.
 V. Nagaraja Ph.D.
V. Nagaraja Ph.D.
Professor, Indian Institute of Science, Bengaluru, India
Perturbation of DNA topology in mycobacteria
To maintain the topological homeostasis of the genome in the cell, DNA topoisomerases catalyse DNA cleavage, strand passage and rejoining of the ends. Thus, although they are essential house- keeping enzymes, they are the most vulnerable targets; arrest of the reaction after the first trans-esterification step leads to breaks in DNA and cell death. Some of the successful antibacterial or anticancer drugs target the step ie arrest the reaction or stabilize the topo -DNA covalent complex. I will describe our efforts in this direction – to target DNA gyrase and also topoisomerase1 from mycobacteria. The latter, although essential, has no inhibitors described so far. The new inhibitors being characterized are also used to probe topoisomerase control of gene expression.
In the biological warfare between the organisms, a diverse set of molecules encoded by invading genomes target the above mentioned most vulnerable step of topoisomerase reaction, leading to the accumulation of double strand breaks. Bacteria, on their part appear to have developed defense strategies to protect the cells from genomic double strand breaks. I will describe a mechanism involving three distinct gyrase interacting proteins which inhibit the enzyme in vitro. However, in vivo all these topology modulators protect DNA gyrase from poisoning effect by sequestering the enzyme away from DNA.
Next, we have targeted a topology modulator protein, a nucleoid associated protein(NAP) from Mycobacterium tuberculosis to develop small molecule inhibitors by structure based design. Over expression of HU leads to alteration in the nucleoid architecture. The crystal structure of the N-terminal half of HU reveals a cleft that accommodates duplex DNA. Based on the structural feature, we have designed inhibitors which bind to the protein and affect its interaction with DNA, de-compact the nucleoid and inhibit cell growth. Chemical probing with the inhibitors reveal the importance of HU regulon in M.tuberculosis.
 R. Manjunath, Ph.D.
R. Manjunath, Ph.D.
Associate Professor, Dept of Biochemistry, Indian Institute of Science, Bengaluru, India
REGULATION OF THE MHC COMPLEX AND HLA SOLUBILISATION BY THE FLAVIVIRUS, JAPANESE ENCEPHALITIS VIRUS
Viral encephalitis caused by Japanese encephalitis virus (JEV) and West Nile Virus (WNV) is a mosquito-borne disease that is prevalent in different parts of India and other parts of South East Asia. JEV is a positive single stranded RNA virus that belongs to the Flavivirus genus of the family Flaviviridae. The genome of JEV is about 11 kb long and codes for a polyprotein which is cleaved by both host and viral encoded proteases to form 3 structural and 7 non-structural proteins. It is a neurotropic virus which infects the central nervous system (CNS) and causes death predominantly in newborn children and young adults. JEV follows a zoonotic life-cycle involving mosquitoes and vertebrate, chiefly pigs and ardeid birds, as amplifying hosts. Humans are infected when bitten by an infected mosquito and are dead end hosts. Its structural, pathological, immunological and epidemiological aspects have been well studied. After entry into the host following a mosquito bite, JEV infection leads to acute peripheral neutrophil leucocytosis in the brain and leads to elevated levels of type I interferon, macrophage-derived chemotactic factor, RANTES,TNF-α and IL-8 in the serum and cerebrospinal fluid.
Major Histocompatibility Complex (MHC) molecules play a very important role in adaptive immune responses. Along with various classical MHC class I molecules, other non-classical MHC class I molecules play an important role in modulating innate immune responses. Our lab has shown the activation of cytotoxic T-cells (CTLs) during JEV infection and CTLs recognize non-self peptides presented on MHC molecules and provide protection by eliminating infected cells. However, along with proinflammatory cytokines such as TNFα, they may also cause immunopathology within the JEV infected brain. Both JEV and WNV, another related flavivirus have been shown to increase MHC class I expression. Infection of human foreskin fibroblast cells (HFF) by WNV results in upregulation of HLA expression. Data from our lab has also shown that JEV infection upregulates classical as well as nonclassical (class Ib) MHC antigen expression on the surface of primary mouse brain astrocytes and mouse embryonic fibroblasts.
There are no reports that have discussed the expression of these molecules on other cells like endothelial and astrocyte that play an important role in viral invasion in humans. We have studied the expression of human classical class I molecules HLA-A, -B, -C and the non-classical HLA molecules, HLA-E as well as HLA-F in immortalized human brain microvascular endothelial cells (HBMEC), human endothelial cell line (ECV304), human glioblastoma cell line (U87MG) and human foreskin fibroblast cells (HFF). Nonclassical MHC molecules such as mouse Qa-1b and its human homologue, HLA-E have been shown to be the ligand for the inhibitory NK receptor, NKG2A/CD94 and may bridge innate and adaptive immune responses. We show that JEV infection of HBMEC and ECV 304 cells upregulates the expression of HLA-A, and –B antigens as well as HLA-E and HLA-F. Increased expression of total HLA-E upon JEV infection was also observed in other human cell lines as well like, human amniotic epithelial cells, AV-3, FL and WISH cells. Further, we show for the first time that soluble HLA-E (sHLA-E) was released from infected ECV and HBMECs. In contrast, HFF cells showed only upregulation of cell-surface HLA-E expression while U87MG, a human glioblastoma cell line neither showed any cell-surface induction nor its solubilization. This shedding of sHLA-E was found to be dependent on matrix metalloproteinase (MMP) and an important MMP, MMP-9 was upregulated during JEV infection. Treatment with IFNγ resulted in the shedding of sHLA-E from ECV as well as U87MG but not from HFF cells. Also, sHLA-E was shed upon treatment with IFNβ and both IFNβ and TNFα, when present together caused an additive increase in the shedding of sHLA-E. HLA-E is an inhibitory ligand for CD94/NKG2A receptor of Natural Killer cells. Thus, MMP mediated solubilization of HLA-E from infected endothelial cells may have important implications in JEV pathogenesis including its ability to compromise the blood brain barrier.
 Sharmila Mande, Ph.D.
Sharmila Mande, Ph.D.
Principal Scientist and Head, Bio Sciences R&D, TCS Innovation Labs, Pune
Gut microbiome and health: Moving towards the new era of translational medicine
The microbes inhabiting our body outnumber our own cells by a factor of 10. The genomes of these microbes, called the ‘second genome’ are therefore expected to have great influence on our health and well being. The emerging field of metagenomics is rapidly becoming the method of choice for studying the microbial community (called microbiomes) present in various parts of the human body. Recent studies have implicated the role of gut microbiomes in several diseases and disorders. Studies have indicated gut microbiome’s role in nutrient absorption, immuno-modulation motor-response, and other key physiological processes. However, our understanding of the role of gut microbiota in malnutrition is currently incomplete. Exploration of these aspects are likely to help in understanding the microbial basis for several physiological disorders associated with malnutrition (eg, increased susceptibility to diarrhoeal pathogens) and may finally aid in devising appropriate probiotic strategies addressing this menace. A metagenomic approach was employed for analysing the differences between gut microbial communities obtained from malnourished and healthy children. Results of the analysis using TCS’ ‘Metagenomic Analysis Platform’ were discussed in detail during my talk.
 Manzoor K, Ph.D.
Manzoor K, Ph.D.
Professor, Centre for Nanoscience & Molecular Medicine, Amrita University
Targeting aberrant cancer kinome using rationally designed nano-polypharmaceutics
Manzoor Koyakutty, Archana Ratnakumary, Parwathy Chandran, Anusha Ashokan, and Shanti Nair
`War on Cancer’ was declared nearly 40 years ago. Since then, we made significant progress on fundamental understanding of cancer and developed novel therapeutics to deal with the most complex disease human race ever faced with. However, even today, cancer remains to be the unconquered `emperor of all maladies’. It is well accepted that meaningful progress in the fight against cancer is possible only with in-depth understanding on the molecular mechanisms that drives its swift and dynamic progression. During the last decade, emerging new technologies such as nanomedicine could offer refreshing life to the `war on cancer’ by way of providing novel methods for molecular diagnosis and therapy.
In the present talk, we discuss our approaches to target critically aberrant cancer kinases using rationally designed polymer-protein and protein-protein core-shell nanomedicines. We have used both genomic and proteomic approaches to identify many intimately cross-linked and complex aberrant protein kinases behind the drug resistance and uncontrolled proliferation of refractory leukemic cells derived from patients. Small molecule inhibitors targeted against oncogenic pathways in these cells were found ineffective due to the involvement of alternative survival pathways. This demands simultaneous inhibition more than one oncogenic kinases using poly-pharmaceutics approach. For this, we have rationally designed core-shell nanomedicines that can deliver several small molecules together for targeting multiple cancer signalling. We have also used combination of small molecules and siRNA for combined gene silencing together with protein kinase inhibition in refractory cancer cells. Optimized nanomedicines were successfully tested in patient samples and found enhanced cytotoxicity and molecular specificity in drug resistant cases.
Nano-polypharmaceutics represents a new generation of nanomedicines that can tackle multiple cancer mechanisms simultaneously. Considering the complexity of the disease, such therapeutic approaches are not simply an advantage, but indispensable.
Acknowledgements:
We thank Dept. of Biotechnology and Dept. Of Science and Technology,Govt. of India for the financial support through `Thematic unit of Excellence in Medical NanoBiotechnology’ and `Nanomedicine- RNAi programs’.
 Andrey Panteleyev, Ph.D.
Andrey Panteleyev, Ph.D.
Vice Chair, Division of Molecular Biology, NBICS Centre-Kurchatov Institute, Moscow, Russia
The system of PAS proteins (HIF and AhR) as an interface between environment and skin homeostasis
Regulation of normal skin functions as well as etiology of many skin diseases are both tightly linked to the environmental impact. Nevertheless, molecular aspects of skin-environment communication and mechanisms coordinating skin response to a plurality of environmental stressors remain poorly understood.
Our studies along with the work of other groups have identified the family of PAS dimeric transcription factors as an essential sensory and regulatory component of communication between skin and the environment. This protein family comprises a number of hypoxia-induced factors (HIF-alpha proteins), aryl hydrocarbon receptor (AhR), AhR nuclear translocator (ARNT), and several proteins implicated in control of rhythmic processes (Clock, Period, and Bmal proteins). Together, various PAS proteins (and first of all ARNT – as the central dimerization partner in the family) control such pivotal aspects of cell physiology as drug/xenobiotic metabolism, hypoxic and UV light response, ROS activity, pathogen defense, overall energy balance and breathing pathways.
In his presentation Dr. Panteleyev will focus on the role of ARNT activity and local hypoxia in control of keratinocyte differentiation and cornification. His recent work revealed that ARNT negatively regulates expression of late differentiation genes through modulation of amphiregulin expression and downstream alterations in activity of EGFR pathway. All these effects are highly dependent on epigenetic mechanisms such as histone deacetylation. Characterisation of hypoxia as a key microenvironmental factor in the skin and the role of HIF pathway in control of dermal vasculature and epidermal functions is another major focus of Dr. Panteleyev’s presentation.
In general, the studies of Dr. Panteleyev’s laboratory provide an insight into the PAS-dependent maintenance of skin homeostasis and point to the potential role of these proteins in pathogenesis of environmentally-modulated skin diseases such as barrier defects, desquamation abnormalities, psoriasis, etc.
 Tim Guilliams, Ph.D.
Tim Guilliams, Ph.D.
Junior Associate Fellow at the Centre for Science and Policy, University of Cambridge
From Camels to Worms: Novel Approaches for Drug Discovery in Parkinson’s Disease
The discovery of novel treatments for neurodegenerative diseases, such as Parkinson’s disease, represents one of the biggest scientific challenges of the 21st century. The development of new tools and models to study the mechanisms underlying neurotoxicity is therefore essential. During my talk, I will outline new strategies for drug design and innovation used during my PhD at the University of Cambridge, which include the combination of fluorescent nematode worms, camelid antibody fragment technology and chemical compounds. These novel approaches will help us to gain insights into the key pathogenic steps involved in Parkinson’s disease and potentially lead to new therapeutic strategies.
 Gokul Das, Ph.D.
Gokul Das, Ph.D.
Co-Director, Breast Disease Site Research Group, Roswell Park Cancer Institute, Buffalo, NY
Probing Estrogen Receptor−Tumor Suppressor p53 Interaction in Cancer: From Basic Research to Clinical Trial
Tumor suppressor p53 and estrogen receptor have opposite roles in the onset and progression of breast cancer. p53 responds to a variety of cellular of stresses by restricting the proliferation and survival of abnormal cells. Estrogen receptor plays an important role in normal mammary gland development and the preservation of adult mammary gland function; however, when deregulated it becomes abnormally pro-proliferative and greatly contributes to breast tumorigenesis. The biological actions of estrogens are mediated by two genetically distinct estrogen receptors (ERs): ER alpha and ER beta. In addition to its expression in several ER alpha-positive breast cancers and normal mammary cells, ER beta is usually present in ER alpha-negative cancers including triple-negative breast cancer. In spite of genetically being wild type, why p53 is functionally debilitated in breast cancer has remained unclear. Our recent finding that ER alpha binds directly to p53 and inhibits its function has provided a novel mechanism for inactivating genetically wild type p53 in human cancer. Using a combination of proliferation and apoptosis assays, RNAi technology, quantitative chromatin immunoprecipitation (qChIP), and quantitative real-time PCR (qRT-PCR), in situ proximity ligation assay (PLA), and protein expression analysis in patient tissue micro array (TMA), we have demonstrated binding of ER alpha to p53 and have delineated the domains on both the proteins necessary for the interaction. Importantly, ionizing radiation inhibits the ER-p53 interaction in vivo both in human cancer cells and human breast tumor xenografts in mice. In addition, antiestrogenstamoxifen and faslodex/fulvestrant (ICI 182780) disrupt the ER-p53 interaction and counteract the repressive effect of ER alpha on p53, whereas 17β-estradiol (E2) enhances the interaction. Intriguingly, E2 has diametrically opposite effects on corepressor recruitment to a p53-target gene promoter versus a prototypic ERE-containing promoter. Thus, we have uncovered a novel mechanism by which estrogen could be providing a strong proliferative advantage to cells by dual mechanisms: enhancing expression of ERE-containing pro-proliferative genes while at the same time inhibiting transcription of p53-dependent anti-proliferative genes. Consistently, ER alpha enhances cell cycle progression and inhibits apoptosis of breast cancer cells. Correlating with these observations, our retrospective clinical study shows that presence of wild type p53 in ER-positive breast tumors is associated with better response to tamoxifen therapy. These data suggest ER alpha-p53 interaction could be one of the mechanisms underlying resistance to tamoxifen therapy, a major clinical challenge encountered in breast cancer patients. We have launched a prospective clinical trial to analyze ER-p53 interaction in breast cancer patient tumors at Roswell Park Cancer Institute. Our more recent finding that ER beta has opposite functions depending on the mutational status of p53 in breast cancer cells is significant in understanding the hard-to-treat triple-negative breast cancer and in developing novel therapeutic strategies against it. Our integrated approach to analyze ER-p53 interaction at the basic, translational, and clinical research levels has major implications in the diagnosis, prognosis, and treatment of breast cancer.
 Shrikant Anant, Ph.D.
Shrikant Anant, Ph.D.
The Department of Molecular & Integrative Physiology, Kansas University Medical Center, USA
Cancer Stem Cells: Target Colon Cancers
Shrikant Anant, Deep Kwatra and Dharmalingam Subramaniam
Colon cancer is a leading cause of cancer related deaths in the US, and its rate is increasing at an alarming rate in lndia. Recent studies have suggested the drug resistance role for a mall number of cells within a tumor called cancer stem cells. We identified the colon cancer stem cell marker DCLK1, a member of the protein kinase superfamily and the doublecortin family. The protein encodes a Cterminal serinethreonine protein kinase domain, which shows substantial homology to Ca2calmodulindependent protein kinase. Our current studies have been to identify compounds that can either affect DCLK1 expression or inhibits its activity as a way to inhibit cancer stem cells. Honokiol is a biphenolic compound that has been used in the traditional Chinese Medicine for treating various ailments. In vitro kinase assays with recombinant DCLK1 demonstrated that honokiol inhibits its kinase activity in a dose dependent manner. We therefore determined the effect of honokiol on stem cells. One method to look at effects on stem cells is perform a spheroid assay, where spheroids formation is suggested to maintain stemlike characteristic of cancer cells. Honokiol significantly suppressed colonosphere formation of two colon cancer cell lines HCT116 and SW480. Flow cytometry studies confirmed that honokiol reduced the number of DCLK1cells. A critical signaling pathway known to modulate intestinal stem cell proliferation is the Hippo signaling pathway, and deregulation of the pathway leads to tumor development. DCLK1cells had high levels of YAP1, the nuclear target of Hippo signaling. We determined the effect of honokiol on components of the hipposignaling pathway. Honokiol reduced the phosphorylation of Mst1/2, Lats1/2 and YAP1. Furthermore, honokiol treatment resulted in downregulation of YAPTEAD complex protein TEAD-1. Ectopic expression of the TEAD-1 partially rescued the cells from honokiol mediated growth suppression. To determine the effect of honokiol on tumor growth in vivo, nude mice harboring HCT116 tumor xenografts in their flanks were administered the compound intraperitoneally every day for 21 days. Honokiol treatment significantly inhibited tumor xenograft growth. Western blot and immunohistochemistry analyses demonstrated significant inhibition in the expression of stem marker and Hippo signaling proteins in the honokioltreated xenograft tissues. Taken together, these data suggest that honokiol is a potent inhibitor of colon cancer that targets DCLK1 stem cells by inhibiting Hippo signaling pathway.
 Bodo Eickhoff, Ph.D.
Bodo Eickhoff, Ph.D.
Senior Vice-President, Head of Sales and Marketing for Roche Applied Science, Germany
New paths for treatment of complex diseases: target combinatorial drug therapy
Several types of diseases show a complex pathogenesis and require targeted as well as combinatorial drug treatment. A classical example, Tuberculosis, was thought for decades to be managable by triple therapy, however now requiring new therapeutic approaches due to multi drug resistant strains. HIV and AIDS can only be kept under control by combinations of specific, virus-protein targeted drugs, requiring constant monitoring of resistance patterns and modulation of drug combinations during life-long therapy. As a third example, Cancer in all its different variations, requires detailled molecular understanding to enable targeted therapy. New technologies provide more and in depths molecular insights into pathomechanisms and resulting treatment options. However, is there an alternative way to approach complex diseases by holistic models? Can restoring of apoptosis-capabilities of transformed cells be an example of such an alternative path? How do we in future adress major unresolved topics like increasing drug resistance in bacterial infections, lack of anti-viral drugs, treatment of parasite diseases like Malaria, and newly emerging infectious diseases in research and fast translation of these results into diagnosis and treatment?
 Rustom Mody, Ph.D.
Rustom Mody, Ph.D.
Head R & D Lupin Ltd., Pune
Biosimilars are follow-on biologics also known as Similar Biologics – terms used to describe officially approved subsequent versions of innovator biopharmaceutical products made by rDNA technology when made by a different sponsor following patent expiry on the innovator product. These products are drawing global attention as a large number of block buster biopharmaceuticals have expired and many will soon seize to have patent protection in the coming years, opening the doors for the entry of biosimilars. However, the regulatory landscape is getting complex across the globe. The talk focuses on opportunities and challenges in the field of biosimilars and the future of biosimilar companies in India.
 Srinivas Pentyala, Ph.D.
Srinivas Pentyala, Ph.D.
Director of Translational Research, Faculty member in Anesthesiology, Urology, Health Sciences, Physiology & Biophysics, Stony Brook Medical Center, Stony Brook, NY, USA
Translational Biotechnology in the development of Drugs, Diagnostics and Devices
Innovation can propel a simple finding in basic science to a concept that can be of immense value to society. While applying these novel innovations that came out by studying the basic mechanisms, an offshoot of applied branch emerged. This field which is now widely referred to as Translational Research utilizes basic science findings and translates these findings using biotech and nanotech principles into innovative concepts for the benefit of mankind. This branch of science has evolved into a multidisciplinary juggernaut encompassing all known fields of science, even including fields as varied as law, economics, sociology, etc. Scientists, who have a passion to commit themselves for the betterment of society, often limit themselves to perform bench research and dwell in the satisfaction of discovering new things. However, this sort of contribution is the nexus for advancement of science. Particularly the field of biotechnology is an area where intense research is being done to understand the physical as well as biological principles ranging from matter to mind. With the ever increasing interest in this branch, and the dreams and aspirations that this field can bring, basic science researchers are now taking a bold step into this new realm, merging different fields of knowledge to come up with novel inventions. This approach though lucrative, needs discipline and a step-by-step systemic approach. Taking basic science discoveries related to diseases and symptoms from our research lab as examples and taking the approach of translational biotechnology, the whole concept of thinking outside the box and taking the next step from basic science to applied and translational sciences for improving the quality of life will be presented. Some of the translational projects that will be discussed are:
- How do studies on a simple calcium mobilizing enzyme help understand and cure diseases like Alzheimer’s?
- The drugs that were routinely given in operating rooms can behave like double edged swords to cure diseases like asthma.
- Using simple antigen – antibody technology, can a novel diagnostic and prognostic tool be discovered for cancer?
- Using the information garnered from the Human Genome project and utilizing the tools of bioinformatics can we develop novel new drugs?
- The concept of delivering drugs and how novel diagnostic tools can be developed to reduce mishaps.
- How do environmental toxins help us to understand complex biological processes like sleep and memory?
- How does biochemistry and bioinformatics help us to have clean drinking water?
The findings from the above projects will be used as examples and different mechanisms of translational research using biotech applications will be discussed.

Manjunath Joshi, Apoorva Lad, Bharat Prasad Alevoor, Aswath Balakrishnan, Lingadakai Ramachandra and Kapaettu Satyamoorthy
Pathological conditions during Type 2 Diabetes (T2D) are associated with elevated risk for common community acquired infections due to poor glycemic control. Multiple studies have indicated specific defects in innate and adaptive immune function in diabetic subjects. Neutrophils play an important role in eliminating pathogens as an active constituent of innate immune system. Apart from canonically known phagocytosis mechanism, neutrophils are endowed with a unique ability to produce extracellular traps (NETs) to kill pathogens by expelling DNA coated with bactericidal proteins and histone. NETosis is stimulated by diverse bacteria and their products, fungi, protozoans, cytokines, phorbol esters and by activated platelets. Considering deregulation of metabolic and immune response pathways during pathological state of diabetes and NETosis as a potential mechanism for killing bacteria, we therefore, investigated whether hyperglycemic conditions modulate formation of neutrophil NETs and attempted to identify underlying immunoregulatory mechanisms. Freshly isolated neutrophils from normal individuals were cultured in absence or presence of high glucose (different concentrations) for 24 hours and activated with either LPS (2 mg/ml) or PMA (20 ng/ml) or IL-6 (20 ng/ml) for 3 hours. NETs were visualized and quantified by addition of DNA binding dye SYTOX green using fluorescence microscope and fluorimetry. NETs were quantified in Normal and diabetic subjects. Serum IL-6 levels were measured using ELISA technique. NETs bound elasatse were quantified in normal and diabetic subjects in presence or absence of DNase. Bacterial killing assays were performed upon infecting E.coli with activated neutrophils from normal and diabetic subjects. Microscopy and fluorimetry analysis suggested dramatic impairment in NETs formation under high glucose conditions. Extracellular DNA lattices formed in hyperglycemic conditions were short lived and unstable leading to rapid disintegration. Subsequent, time course experiments showed that NETs production was delayed in hyperglycemic conditions. To validate our findings more closely to clinical conditions, we investigated the neutrophil activation and NETs formation in diabetic patients. Upon stimulation with LPS for three hours, neutrophils from diabetic subjects responded weakly to LPS and lesser NETs were formed; whereas, neutrophils from normal individuals showed robust release of NETs. In few patients we found short and imperfect NETs in basal conditions suggesting constitutive activation of neutrophils in diabetic subjects. Interestingly, NETs bound elastase activity was reduced in diabetes subjects when compared to non-diabetic individuals, indicating a dysfunction of one of the important protein component of NETs during diabetes. Neutrophils from diabetic subjects released higher levels of IL-6 without any stimulation suggesting an existence of constitutively activated pro-inflammatory state. IL-6 induced NETs formation and was abrogated by high glucose. Weobserved that glycolysis inhibitor 2-DG resensitize the high glucose attenuated LPS and IL-6 induced NETs. a) NETs are influenced by glucose homeostasis, b) IL-6 as potent inducer of energy dependent NETs formation and c) hyperglycemia mimics a state of constitutively active pro-inflammatory condition in neutrophils leading to reduced response to external stimuli making diabetic subjects susceptible for infections.

Tanu Sharma, Narendra Parihar, Someshwar Nath, Azad Singh, Savneet Kaur, Simendra Singh and Chandi Mandal.
In the recent years, there has been a surge in various forms of cancers especially in urban areas of India. Tumor recurrence and metastasis are the main causes of increased morbidity and mortality. In poor and developing countries like India chemotherapy is the choosen method of therapeutic intervention for different types of malignancies such as breast, prostate, pancreas, etc. Although general chemotherapy treatment is effective in controlling tumour growth but high doses of the treatment regimen often results in severe off-target toxicity to different organs including liver, kidney, heart, brain etc. (1) A large percentage of patients are unable to tolerate this toxicity and sometime it can also lead to life threatening complications. (2) Therefore, the primary objective of this study is to prevent this druginduced off-target toxicity. Omega 3 fatty acids (n-3FAs) such as DHA (docosahexaenoic acid) and EPA (eicosapentaenoic acid) that are active components of fish oil play active role in preventing cancer growth and its metastasis. (3–5) We have recently identified that omega 3-fatty acids prevent breast cancer bone metastasis by targeting cancer stem cell marker CD44 and osteoclastogenic factor CSF-1(4, 5). With this background, the present study addresses the role of n-3FAs in preventing chemotherapeutic drug-induced off-target toxicity.
Methods
The cat fishes were divided into four groups (each group contains 4 fishes) for each set of experiments. Three different chemotherapeutic drugs etoposide, doxorubicin, cisplatin at higher dosages (4, 2, 2 mg/kg body wt respectively) and fish oil (60mg/kg body wt, DHA: EPA; 2:3) were administered to cat fish (Mangur) by gavage. After 6 days, cat fishes were sacrificed and different organs were isolated. SGPT and alkaline phosphatase (ALP) activity assays were performed to examine liver function and superoxide dismutase activity (SOD) assay was conducted to evaluate reactive oxygen species (ROS) level.
Results
Severe skin damages were observed both in doxorubicin and cisplatintreated cat fishes as compared to the control fishes, indicating side effects of chemotherapeutic drugs treatment. High level of SGPT as well as ALP activity was observed in liver samples of etoposide and doxorubicin – treated fishes, suggesting chemotherapeutic drug-induced liver toxicity. Mechanistically, we found that doxorubicin treatment showed significant decrease of SOD activity in liver samples in comparison to the control fishes, suggesting that chemotherapeutic drugs lead to organ toxicity presumably by increasing ROS levels (6). However, treatment of the fishes with n-3FAs of the fish oil led to a dramatic reduction of chemotherapeutic drug-induced skin damages. Also, low levels of SGPT and ALP activities were observed in the fishes given etoposide in combination with n-3FAs as compared to the fishes given etoposide alone. Similarly, fish oil also gave protection against doxorubicin-induced liver dysfunction. Our data further showed that n-3FAs treatment significantly increased etoposide and doxorubicin-inhibited SOD activity.
Conclusions
The study for the first time reports that the use of n-3FAs leads to a dramatic reduction of chemotherapeutic drug-induced skin damages and mitigates chemotherapeutic drug-driven liver dysfunction presumably by reducing ROS level. This study suggests that n-3FAs may possibly be used in combination with chemotherapeutic drugs to treat different cancers to reduce chemotherapeutic drugs-associated systemic toxicity and to increase anticancer activity. Ongoing research study will further address the role of n-3FAs on other off-target toxic effects of chemotherapeutic drugs.

Aswath Balakrishnan, Kapaettu Satyamoorthy and Manjunath B Joshi
Introduction
Insulin resistance is a hall mark of metabolic disorders such as diabetes. Reduced insulin response in vasculature leads to disruption of IR/Akt/eNOS signaling pathway resulting in vasoconstriction and subsequently to cardiovascular diseases. Recent studies have demonstrated that inflammatory regulator interleukin-6 (IL-6), as one of the potential mediators that can link chronic inflammation with insulin resistance. Accumulating evidences suggest a significant role of epigenetic mechanisms such as DNA methylation in progression of metabolic disorders. Hence the present study aimed to understand the role of epigenetic mechanisms involved during IL-6 induced vascular insulin resistance and its consequences in cardiovascular diseases.
Materials and Methods
Human umbilical vein endothelial cells (HUVEC) and Human dermal microvascular endothelial cells (HDMEC) were used for this study. Endothelial cells were treated in presence or absence of IL-6 (20ng/ml) for 36 hours and followed by insulin (100nM) stimulation for 15 minutes. Using immunoblotting, cell lysates were stained for phosphor- and total Akt levels to measure insulin resistance. To investigate changes in DNA methylation, cells were treated with or without neutrophil conditioned medium (NCM) as a physiological source of inflammation or IL-6 (at various concentrations) for 36 hours. Genomic DNA was processed for HPLC analysis for methyl cytosine content and cell lysates were analyzed for DNMT1 (DNA (cytosine-5)-methyltransferase 1) and DNMT3A (DNA (cytosine-5)-methyltransferase 3A) levels using immunoblotting.
Results
Endothelial cells stimulated with insulin exhibited an increase in phosphorylation of Aktser 473 in serum free conditions but such insulin response was not observed in cells treated with IL-6, suggesting chronic exposure of endothelial cells to IL-6 leads to insulin resistance. HPLC analysis for global DNA methylation resulted in decreased levels of 5-methyl cytosine in cells treated with pro-inflammatory molecules (both by NCM and IL-6) as compared to untreated controls. Subsequently, analysis in cells treated with IL-6 showed a significant decrease in DNMT1 levels but not in DNMT3A. Other pro-inflammatory marker such as TNF-α did not exhibit such changes.
Conclusion
Our study suggests: a) Chronic treatment of endothelial cells with IL-6 results in insulin resistance b) Neutrophil conditioned medium and IL-6 decreases methyl cytosine levels c) DNMT1 but not DNMT3a levels are reduced in cells treated with IL-6.

Sunilkumar Sukumaran, Ayyappan Nair, Madhuri Subbiah, Gunja Gupta, Lakshmi Rajakrishna, Pradeep Savanoor Raghavendra, Subbulakshmi Karthikeyan, Salini Krishnan Unni and Ganesh Sambasivam
Genotoxicity is defined as DNA damage that leads to gene mutations which can become tumorigenic. Genotoxicity testing is important to ensure drug safety and is mandatory prior to Phase I/II clinical trials of new drugs. The results from genetic toxicology studies help to identify hazardous drugs and environmental genotoxins. Currently, among others there are four tests recommended by regulatory authorities (Ames test-bacterial, chromosome aberrations; in vitro gene mutation-eukaryotic cells and in vivo test). These assays are laborious, time consuming, require large quantities of test compounds and limited by throughput challenges. The site and mechanism of genotoxicity are not revealed by these assays and data obtained from bacterial tests might not translate the same in mammals. To address these we have developed a novel, versatile, human cell based, high throughput, reporter based genotoxicity screen (Anthem’s Genotox screen). This screen is performed on genetically engineered human cell lines that express 3 reporter genes under transcriptional control of ‘early DNA damage sensors’ (p53, p21 and GADD153). These genes are involved in DNA repair, cell cycle arrest and/or apoptosis. p21 and GADD are also known to be induced in a p53 independent manner. p53 blocks G1/S transition of cell cycle while the p53 independent DNA damage block G2/M transition. Identification of the mechanism of genotoxicity helps in rational drug designing. Additionally, the platform can be used to screen other potential genotoxins from cosmetics, food and environment. Initial validation studies of the Genotox screen was performed with over 60 compounds chosen from a variety of chemical classes. The genotoxic potential of metabolites was tested using rat liver S9 fractions. The results demonstrated a sensitivity of 86.7–92.3% and a specificity of 70–78.6% when compared with currently available in vitro genotoxicity assays. This Genotox screen would prove to be an invaluable human cell based tool to weed out potential genotoxins in various industries.

Jaipal Panga Reddy, Dulal Panda and Sanjeeva Srivastava
The rapid emergence of microbial resistance against the ever increasing number of infectious diseases has been a major hurdle in hunt for novel antibiotics and inventive targets required to combat these dreaded pathogens. Despite the discovery of synthetic and semi-synthetic drugs, infectious diseases remain a major cause of concern. The process of drug discovery started from natural products as an antibacterial drug, an old art which was widely adopted in ancient civilizations in India and China. Most of the existing antibiotics are derived from the backbone of natural compounds (Butler M S 2005). Drug discovery process from natural products to synthetic medicine has continued for thousands of years to battle with pathogenic organisms. Although synthetic drugs played a vital role as antimicrobial drugs during the last decade, the over-use of antibiotics has led to the unanticipated changes at the genomic level, resulting into emergence of antibiotic-resistant strains (Singh P et al. 2010). To combat drug-resistant strains there is a need to develop and characterize new drugs by screening natural and synthetic compounds. Curcumin is a well known natural product, with a potential to cure wide range of cancers. Apart from antitumor activity, it also has anti-inflammatory, anti-viral, anti-genotoxic, phototoxic and antimicrobial activities. The exact mechanism of action of curcumin is still obscure and further investigations are required to identify its molecular/cellular targets. Recently, it was observed that curcumin is able to perturb the FtsZ assembly dynamics and elongate the bacterial cell length by inhibiting bacterial cell division (Rai D et al. 2008).
In the present study, we aimed at deciphering the mechanism of action and molecular targets of curcumin in B. subtilis AH75 using classical two dimensional electrophoresis, 2D-difference gel electrophoresis (2DDIGE) in combination with MALDI-TOF/TOFMS. Comparative proteome analysis of control and IC50 curcumin treated B. subtilis has revealed differential expression of 48 proteins (p ≤ 0.05) in response to curcumin treatment. Further, in silico analysis has revealed the involvement of the differential expressed proteins in protein synthesis, transcription/nucleotide synthesis, stress regulation, central metabolism and amino acid metabolic process. Additionally, suppression in major central metabolic dehydrogenase such as glyceraldehyde-3-phosphate dehydrogenase 2, dihydrolipoyl dehydrogenase, malate dehydrogenase, pyruvate dehydrogenase E1 component subunit beta, 2-oxoglutarate dehydrogenase E1 component, ATP synthase epsilon chain, ATP synthase subunit beta and probable NADdependent malic enzyme 1 has been identified (Figure 1). Reduction in expression levels of major dehydrogenases indicates the disturbance in cell membrane integrity to transfer the electrons from NADH to molecular oxygen or maintain the PMF or maintain the membrane potential. Selected potential proteins obtained from proteomic analysis were validated with real-time expression analysis and metabolic assays such as resazurin assay for metabolic activity, CTC assay for respiratory activity and propidium iodide staining for membrane integrity. Findings from this study have revealed that curcumin has potential effects on multiple physiological pathways in bacteria and inhibition of the cell division machinery is one of the prime targets of this drug. Multiple proteins involved in cell division process such as GroEL, ClpC, MetK, Tuf and major dehydrogenases are significantly modulated as a consequence of the drug treatment.