 Krishnakumar Menon, Ph.D.
Krishnakumar Menon, Ph.D.
Associate Professor, Centre for Nanosciences & Molecular Medicine, Amrita University, Kochi, India
A Far-Western Clinical Proteomics Approach to Detect Molecules of Clinical and Pathological Significance in the Neurodegenerative Disease Multiple Sclerosis.
Multiple Sclerosis (MS), an autoimmune neurodegenerative disorder of the central nervous system. The disease affects young adults at their prime age leading to severe debilitation over several years. Despite advances in MS research, the cause of the disease remains elusive. Thus, our objective is to identify novel molecules of pathological and diagnostic significance important in the understanding, early diagnosis and treatment of MS. Biological fluids such as cerebrospinal fluid (CSF), that bathe the brain serve as a potential source for the identification of pathologically significant autoantibody reactivity in MS. In this regard, we report the development of an unbiased clinical proteomics approach for the detection of reactive CSF molecules that target brain proteins from patients with MS. Proteins of myelin and myelin-axolemmal complexes were separated by two-dimensional gel electrophoresis, blotted onto membranes and probed separately with biotinylated unprocessed CSF samples. Protein spots that reacted specifically to MS-CSF’s were further analyzed by matrix assisted laser desorption ionization-time-of-flight time-of-flight mass spectrometry. In addition to previously reported proteins found in MS, we have identified several additional molecules involved in mitochondrial and energy metabolism, myelin gene expression and/or cytoskeletal organization. Among these identified molecules, the cellular expression pattern of collapsin response mediator protein-2 and ubiquitin carboxy-terminal hydrolase L1 were investigated in human chronic-active MS lesions by immunohistochemistry. The observation that in multiple sclerosis lesions phosphorylated collapsin response mediator protein-2 was increased, whereas Ubiquitin carboxy-terminal hydrolase L1 was down-regulated, not only highlights the importance of these molecules in the pathology of this disease, but also illustrates the use of our approach in attempting to decipher the complex pathological processes leading to multiple sclerosis and other neurodegenerative diseases. Further, we show that in clinicaly isolated syndrome (CIS), we could identify important molecules that could serve as an early diagnostic biomarker in MS.
 Claudia AM Wheeler-Kingshott, Ph.D.
Claudia AM Wheeler-Kingshott, Ph.D.
University Reader in Magnetic Resonance Physics, Department of Neuroinflammation, UCL Institute of Neurology, London, UK
Abstract
Detecting neuronal activity in vivo non-invasively is possible with a number of techniques. Amongst these, in 1990 functional magnetic resonance imaging (fMRI) was proposed as a technique that has a great ability to spatially map brain activity by exploiting the blood oxygenation level dependent (BOLD) contrast mechanism [1, 2]. In fact, neuronal activation triggers a demand for oxygen and induces a localised increase in blood flow and blood volume, which actually exceeds the metabolic needs. This in turns causes an increase of oxyhaemoglobin in the venous compartment, which is a transient phenomenon and is accompanied by a transient change (decrease) in the concentration of deoxyhaemoglobin. Due to its paramagnetic properties, the amount of deoxyhaemoglobin present in the venous blood affects the local magnetic field seen by the spins (protons) and determines the local properties of the MR signal. A decrease in deoxyhaemoglobin during neuronal activity, therefore, induces local variations of this magnetic field that increases the average transverse relaxation time of tissue, measured via the T2* parameter [3]. This means that there is an increase of the MR signal (of the order of a few %, typically <5%) linked to metabolic changes happening during brain function. Activation can be inferred at different brain locations by performing tasks while acquiring the MR signal and comparing periods of rest to periods of activity.
The macroscopic changes of the BOLD signal are well characterised, while the reason for the increased blood supply, exceeding demands, needs further thoughts. Here we will discuss two approaches for explaining the BOLD phenomenon, one that links it to adenosine triphosphate production [4] and enzyme saturation, the other that relates it to the very slow diffusion of oxygen through the blood-brain-barrier with a consequent compensatory high demand of oxygen [5]. Some evidence of restricted oxygen diffusion has been shown by means of hypercapnia [6], although it is not excluded that both mechanisms may be present.
Overall, the BOLD signal changes theory and its physiological basis will be presented and discussed.
References
- Ogawa, S., et al., Brain magnetic resonance imaging with contrast dependent on blood oxygenation. Proc Natl Acad Sci U S A, 1990. 87(24): p. 9868-72.
- Kwong, K.K., et al., Dynamic magnetic resonance imaging of human brain activity during primary sensory stimulation. Proc Natl Acad Sci U S A, 1992. 89(12): p. 5675-9.
- Bandettini PA, et al. Spin-echo and gradient-echo EPI of human brain activation using BOLD contrast: a comparative study at 1.5 T. NMR Biomed. 1994 Mar;7(1-2):12-20
- Fox, P.T., et al., Nonoxidative glucose consumption during focal physiologic neural activity. Science, 1988. 241(4864): p. 462-4.
- Gjedde, A., et al. Reduction of functional capillary density in human brain after stroke. J Cereb Blood Flow Metab, 1990. 10(3): p. 317-26.
- Hoge, R.D., et al., Linear coupling between cerebral blood flow and oxygen consumption in activated human cortex. Proc Natl Acad Sci U S A, 1999. 96(16): p. 9403-8.
 K. Satyamoorthy, Ph.D.
K. Satyamoorthy, Ph.D.
Director, Life Sciences Centre, Manipal University, India
Epigenetic Changes due to DNA Methylation in Human Epithelial Tumors
Extensive global hypomethylation in the genome and hypermthylation of selective tumor specific suppressor genes appears to be a hallmark of human cancers. Data suggests that hypermethylation of promoter region in genes is more closely related to subsequent gene expression; contrary to gene-body DNA methylation. The intricate balance between these two may contribute to the progressive process of development, differentiation and carcinogenesis. Epigenetic changes encompass, apart from DNA methylation, chromatin modifications through post-translational changes in histones and control by miRNAs. At the genome level, effects from these are compounded by copy number variations (CNVs) which may ultimately influence protein functions. From clinical perspective, changes in DNA methylation occur very early which are reversible and are influenced by environmental factors. Therefore, these can be potential resource for identifying therapeutic targets as well as biomarkers for early screening of cancer. Our current efforts in profiling genome wide DNA methylation changes in oral, cervical and breast cancers through DNA methylation microarray analysis has revealed number of alterations critical for survival, progression and metastatic behavior of tumors. Bioinformatics and functional analysis revealed several key regulatory molecules controlled by DNA methylation and suggests that DNA methylation changes in several CpG islands appear to co-segregate in the regions of miRNAs as well as in the CNVs. We have validated the signatures for methylation of CpG islands through bisufite sequencing for essential genes in clinical samples and have undertaken transcriptional and functional analysis in tumor cell lines. These results will be presented.
 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
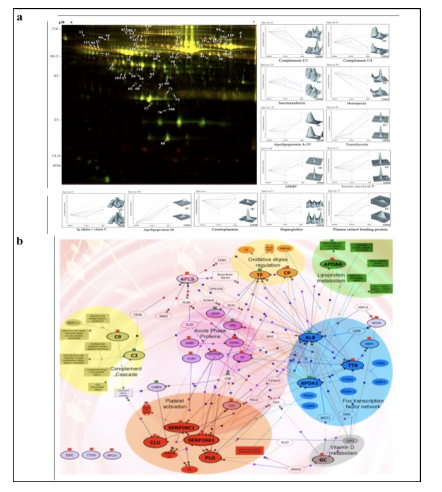
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)

Jaydeep Unni, Ph.D.
Sr. Project Manager, Robert Bosch Healthcare Systems, Palo Alto, CA
Remote Patient Monitoring – Challenges and Opportunities
Remote Patient Monitoring (RPM) is gaining importance and acceptance with rising number of chronic disease conditions and with increase in the aging population. As instances of Heart diseases, Diabetes etc are increasing the demand for these technologies are increasing. RPM devices typically collect patient vital sign data and in some case also patient responses to health related questions. Thus collected data is then transmitted through various modalities (wireless/Bluetooth/cellular) to Hospitals/Doctor’s office for clinical evaluation. With these solutions Doctors are able to access patient’s vital data ‘any time any where’ thus enabling them to intervene on a timely and effective manner. For older adult population chronic disease management, post-acute care management and safety monitoring are areas were RPM finds application. That said, there are significant challenges in adoption of Remote Patient Monitoring including patient willingness and compliance for adoption, affordability, availability of simpler/smarter technology to mention a few. But experts contend that if implemented correctly Remote Patient Monitoring can contain healthcare expenditure by reducing avoidable hospitalization while greatly improving quality of care.
 Kal Ramnarayan, Ph.D.
Kal Ramnarayan, Ph.D.
Co-founder President & Chief Scientific Officer, Sapient Discovery, San Diego, CA, USA
A cost-effective approach to Protein Structure-guided Drug Discovery: Aided by Bioinformatics, Chemoinformatics and computational chemistry
With the mapping of the human genome completed almost a decade ago, efforts are still underway to understand the gene products (i.e., proteins) in the human biological and disease pathways. Deciphering such information is very important for the discovery and development of small molecule drugs as well as protein therapeutics for various human diseases for which no cure exists. As an example, with more than 500 members, the kinase family of protein targets continues to be an important and attractive class for drug discovery. While how many of the members in this family are actually druggable is still to be established, there are several ongoing efforts on this class of proteins across a broad spectrum of disease categories. Even though in general the protein structural topology might looks similar, there are issues with respect selectivity of identified small molecule inhibitors when, the lead molecule discovery is carried out at the ATP binding site. As an added complexity, allosteric modulators are needed for some of the members, but the actual site for such modulation on the protein target can not resolved with uncertainty. In this presentation we will describe a bioinformatics and computational based platform for small molecule discovery for protein targets that are involved in protein-protein interactions as well as targets like kinases and phosphatases. We will describe a computational approach in which we have used an informatics based platform with several hundred kinases to sort through in silico and identify inhibitors that are likely to be highly selective in the lead generation phase. We will discuss the implication of this approach on the drug discovery of the kinase and phosphatase classes in general and independent of the disease category.

Tejaswini Subbannayya, Nandini A. Sahasrabuddhe, Arivusudar Marimuthu, Santosh Renuse, Gajanan Sathe, Srinivas M. Srikanth, Mustafa A. Barbhuiya, Bipin Nair, Juan Carlos Roa, Rafael Guerrero-Preston, H. C. Harsha, David Sidransky, Akhilesh Pandey, T. S. Keshava Prasad and Aditi Chatterjee
Proteomic profiling of gallbladder cancer secretome – a source for circulatory biomarker discovery
Gallbladder cancer (GBC) is the fifth most common cancer of the gastrointestinal tract and one of the common malignancies that occur in the biliary tract (Misra et al. 2006; Lazcano-Ponce et al. 2001). It has a poor prognosis with survival of less than 5 years in 90% of the cases (Misra et al. 2003). The etiology is ill-defined. Several risk factors have been reported including cholelithiasis, obesity, female gender and exposure to carcinogens (Eslick 2010; Kumar et al. 2006). Poor prognosis in GBC is mainly due to late presentation of the disease and lack of reliable biomarkers for early diagnosis. This emphasizes the need to identify and characterize cancer biomarkers to aid in the diagnosis and prognosis of GBC. Secreted proteins are an important class of molecules which can be detected in body fluids and has been targeted for biomarker discovery. There are challenges faced in the proteomic interrogation of body fluids especially plasma such as low abundance of tumor secreted proteins, high complexity and high abundance of other proteins that are not released by the tumor cells (Tonack et al. 2009). Profiling of conditioned media from the cancer cell lines can be used as an alternate means to identify secreted proteins from tumor cells (Kashyap et al. 2010; Marimuthu et al. 2012). We analyzed the invasive property of 7 GBC cell lines (SNU-308, G-415, GB-d1, TGBC2TKB, TGBC24TKB, OCUG-1 and NOZ). Four cell lines were selected for analysis of the cancer secretome based on the invasive property of the cells. We employed isobaric tags for relative and absolute quantitation (iTRAQ) labeling technology coupled with high resolution mass spectrometry to identify and characterize secretome from the panel of 4GBC cancer cells mentioned above. In total, we have identified around 2,000 proteins of which 175 were secreted at differential abundance across all the four cell lines. This secretome analysis will act as a reservoir of candidate biomarkers. Currently, we are investigating and validating these candidate markers from GBC cell secretome. Through this study, we have shown mass spectrometry-based quantitative proteomic analysis as a robust approach to investigate secreted proteins in cancer cells.



