 K. Satyamoorthy, Ph.D.
K. Satyamoorthy, Ph.D.
Director, Life Sciences Centre, Manipal University, India
Epigenetic Changes due to DNA Methylation in Human Epithelial Tumors
Extensive global hypomethylation in the genome and hypermthylation of selective tumor specific suppressor genes appears to be a hallmark of human cancers. Data suggests that hypermethylation of promoter region in genes is more closely related to subsequent gene expression; contrary to gene-body DNA methylation. The intricate balance between these two may contribute to the progressive process of development, differentiation and carcinogenesis. Epigenetic changes encompass, apart from DNA methylation, chromatin modifications through post-translational changes in histones and control by miRNAs. At the genome level, effects from these are compounded by copy number variations (CNVs) which may ultimately influence protein functions. From clinical perspective, changes in DNA methylation occur very early which are reversible and are influenced by environmental factors. Therefore, these can be potential resource for identifying therapeutic targets as well as biomarkers for early screening of cancer. Our current efforts in profiling genome wide DNA methylation changes in oral, cervical and breast cancers through DNA methylation microarray analysis has revealed number of alterations critical for survival, progression and metastatic behavior of tumors. Bioinformatics and functional analysis revealed several key regulatory molecules controlled by DNA methylation and suggests that DNA methylation changes in several CpG islands appear to co-segregate in the regions of miRNAs as well as in the CNVs. We have validated the signatures for methylation of CpG islands through bisufite sequencing for essential genes in clinical samples and have undertaken transcriptional and functional analysis in tumor cell lines. These results will be presented.
 Deepthy Menon, Ph.D.
Deepthy Menon, Ph.D.
Associate Professor, Centre for Nanosciences & Molecular Medicine, Health Sciences Campus, Amrita University, Kochi, India
Nanobioengineering of implant materials for improved cellular response and activity
Deepthy Menon, Divyarani V V, Chandini C Mohan, Manitha B Nair, Krishnaprasad C & Shantikumar V Nair
Abstract
Current trends in biomaterials research and development include the use of surfaces with topographical features at the nanoscale (dimensions < 100 nm), which influence biomolecular or cellular level reactions in vitro and in vivo. Progress in nanotechnology now makes it possible to precisely design and modulate the surface properties of materials used for various applications in medicine at the nanoscale. Nanoengineered surfaces, owing to their close resemblance with extracellular matrix, possess the unique capacity to directly affect protein adsorption that ultimately modulates the cellular adhesion and proliferation at the site of implantation. Taking advantage of this exceptional ability, we have nanoengineered metallic surfaces of Titanium (Ti) and its alloys (Nitinol -NiTi), as well as Stainless Steel (SS) by a simple hydrothermal method for generating non-periodic, homogeneous nanostructures. The bio- and hemocompatibility of these nanotextured metallic surfaces suggest their potential use for orthopedic, dental or vascular implants. The applicability of nanotextured Ti implants for orthopedic use was demonstrated in vivo in rat models, wherein early-stage bone formation at the tissue-implant interface without any fibrous tissue intervention was achieved. This nanoscale topography also was found to critically influence bacterial adhesion in vitro, with decreased adherence of staphylococcus aureus. The same surface nanotopography also was found to provide enhanced proliferation and functionality of vascular endothelial cells, suggesting its prospective use for developing an antithrombotic stent surface for coronary applications. Clinical SS & NiTi stents were also modified based on this strategy, which would offer a suitable solution to reduce the probability of late stent thrombosis associated with bare metallic stents. Thus, we demonstrate that nanotopography on implant surfaces has a critical influence on the fate of cells, which in turn dictates the long term success of the implant.
Acknowledgement: Authors gratefully acknowledge the financial support from Department of Biotechnology, Government of India through the Bioengineering program.
 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
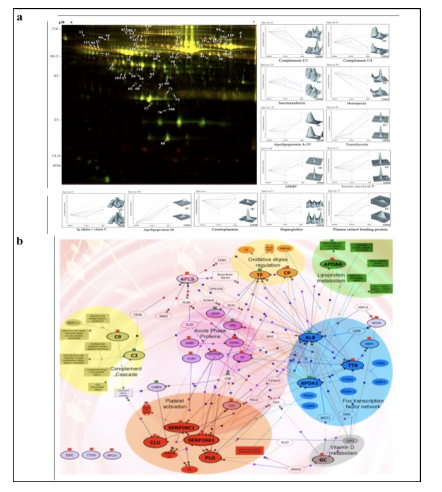
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)
 Michelle Hermiston, MD, Ph.D.
Michelle Hermiston, MD, Ph.D.
Assistant Professor, Department of Pediatrics University of California San Francisco, USA
Interrogating Signaling Networks at the Single Cell Level In Primary Human Patient Samples
Multiparameter phosphoflow cytometry is a highly sensitive proteomic approach that enables monitoring of biochemical perturbations at the single cell level. By combining antisera to cell surface markers and key intracellular proteins, perturbations in signaling networks, cell survival and apoptosis mediators, cell cycle regulators, and/or modulators of other cellular processes can be analyzed in a highly reproducible and sensitive manner in the basal state and in response to stimulation or drug treatment. Advantages of this approach include the ability to identify the biochemical consequences of genetic and/or epigenetic changes in small numbers of cells, to map potential interplay between various signaling networks simultaneously in a single cell, and to interrogate potential mechanisms of drug resistance or response in a primary patient sample. Application of this technology to patients with acute lymphoblastic leukemia or the autoimmune disease systemic lupus erythematosus (SLE) will be discussed.
 Shrikant Anant, Ph.D.
Shrikant Anant, Ph.D.
The Department of Molecular & Integrative Physiology, Kansas University Medical Center, USA
Cancer Stem Cells: Target Colon Cancers
Shrikant Anant, Deep Kwatra and Dharmalingam Subramaniam
Colon cancer is a leading cause of cancer related deaths in the US, and its rate is increasing at an alarming rate in lndia. Recent studies have suggested the drug resistance role for a mall number of cells within a tumor called cancer stem cells. We identified the colon cancer stem cell marker DCLK1, a member of the protein kinase superfamily and the doublecortin family. The protein encodes a Cterminal serinethreonine protein kinase domain, which shows substantial homology to Ca2calmodulindependent protein kinase. Our current studies have been to identify compounds that can either affect DCLK1 expression or inhibits its activity as a way to inhibit cancer stem cells. Honokiol is a biphenolic compound that has been used in the traditional Chinese Medicine for treating various ailments. In vitro kinase assays with recombinant DCLK1 demonstrated that honokiol inhibits its kinase activity in a dose dependent manner. We therefore determined the effect of honokiol on stem cells. One method to look at effects on stem cells is perform a spheroid assay, where spheroids formation is suggested to maintain stemlike characteristic of cancer cells. Honokiol significantly suppressed colonosphere formation of two colon cancer cell lines HCT116 and SW480. Flow cytometry studies confirmed that honokiol reduced the number of DCLK1cells. A critical signaling pathway known to modulate intestinal stem cell proliferation is the Hippo signaling pathway, and deregulation of the pathway leads to tumor development. DCLK1cells had high levels of YAP1, the nuclear target of Hippo signaling. We determined the effect of honokiol on components of the hipposignaling pathway. Honokiol reduced the phosphorylation of Mst1/2, Lats1/2 and YAP1. Furthermore, honokiol treatment resulted in downregulation of YAPTEAD complex protein TEAD-1. Ectopic expression of the TEAD-1 partially rescued the cells from honokiol mediated growth suppression. To determine the effect of honokiol on tumor growth in vivo, nude mice harboring HCT116 tumor xenografts in their flanks were administered the compound intraperitoneally every day for 21 days. Honokiol treatment significantly inhibited tumor xenograft growth. Western blot and immunohistochemistry analyses demonstrated significant inhibition in the expression of stem marker and Hippo signaling proteins in the honokioltreated xenograft tissues. Taken together, these data suggest that honokiol is a potent inhibitor of colon cancer that targets DCLK1 stem cells by inhibiting Hippo signaling pathway.





