 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
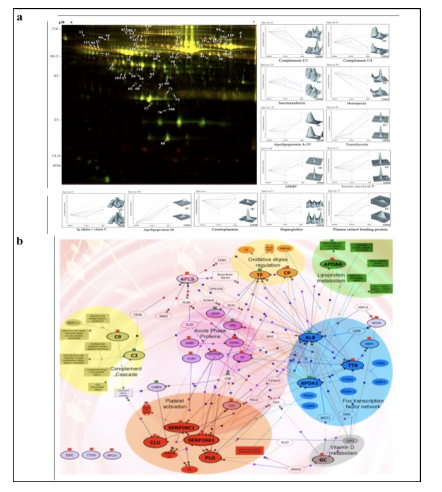
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)
 K. P. Mohanakumar, Ph.D.
K. P. Mohanakumar, Ph.D.
Chief Scientist, Cell Biology & Physiology Division, Indian Institute of Chemical Biology, Kolkata
Neuroprotective and neurodestructive effects of Ayurvedic drug constituents: Parkinson’s disease
The present study reports the good and the bad entities in an Indian traditional medicine used for treating Parkinson’s disease (PD). A prospective clinical trial on the effectiveness of Ayurvedic medication in a population of PD patients revealed significant benefits, which has been attributed to L-DOPA present in the herbs [1]. Later studies revealed better benefits with one of the herbs alone, compared to pure L-DOPA in a clinical trial conducted in UK [2], and in several studies conducted on animal models of PD in independent laboratories world over [3-5]. We have adapted strategies to segregate molecules from the herb, and then carefully removed L-DOPA contained therein, and tested each of these sub-fractions for anti-PD activity in 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine, rotenone and 6-hydroxydopamine -induced parkinsonian animal models, and transgenic mitochondrial cybrids. We report here two classes of molecules contained in the herb, one of which possessed severe pro-parkinsonian (phenolic amine derivatives) and the other having excellent anti-parkinsonian potential (substituted tetrahydroisoquinoline derivatives). The former has been shown to cause severe dopamine depletion in the striatum of rodents, when administered acutely or chronically. It also caused significant behavioral aberrations, leading to anxiety and depression [6]. The latter class of molecules administered in PD animal model [7], caused reversal of behavioral dysfunctions and significant attenuation of striatal dopamine loss. These effects were comparable or better than the effects of the anti-PD drugs, selegiline or L-DOPA. The mechanism of action of the molecule has been found to be novel, at the postsynaptic receptor signaling level, as well as cellular α-synuclein oligomerization and specifically at mitochondria. The molecule helped in maintaining mitochondrial ETC complex activity and stabilized cellular respiration, and mitochondrial fusion-fission machinery with specific effect on the dynamin related protein 1. Although there existed significant medical benefits that could be derived to patients due to the synergistic actions of several molecules present in a traditional preparation, accumulated data in our hands suggest complicated mechanisms of actions of Ayurvedic medication. Our results also provide great hope for extracting, synthesizing and optimizing the activity of anti-parkinsonian molecules present in traditional Ayurvedic herbs, and for designing novel drugs with novel mechanisms of action.
- N, Nagashayana, P Sankarankutty, MRV Nampoothiri, PK Mohan and KP Mohanakumar, J Neurol Sci. 176, 124-7, 2000.
- Katzenschlager R, Evans A, Manson A, Patsalos PN, Ratnaraj N, Watt H, Timmermann L, Van der Giessen R, Lees AJ. J Neurol Neurosurg Psychiatry.75, 1672-7, 2004.
- Manyam BV, Dhanasekaran M, Hare TA. Phytother Res. 18, 706-12, 2004.
- Kasture S, Pontis S, Pinna A, Schintu N, Spina L, Longoni R, Simola N, Ballero M, Morelli M. Neurotox Res. 15, 111-22, 2009.
- Lieu CA, Kunselman AR, Manyam BV, Venkiteswaran K, Subramanian T. Parkinsonism Relat Disord.16, 458-65, 2010.
- T Sengupta and KP Mohanakumar, Neurochem Int. 57, 637-46, 2010.
- T Sengupta, J Vinayagam, N Nagashayana, B Gowda, P Jaisankar and KP Mohanakumar, Neurochem Res 36, 177-86, 2011

David Ibanez, Laura Dubreuil and Alejandro Rier
Neurofeedback (NF) is a type of biofeedback that uses real time display of electroencephalography to illustrate brain activity. EEG features are extracted and displayed allowing the user to, with practice, modulate their temporal evolution. Neurofeedback training has many therapeutic applications such as attention deficit hyperactivity disorder (ADHD), migraine, depression or conduct disorders. This document presents NeuroSurfer, a novel general-purpose tool for neurofeedback training with a use case of attention deficit hyperactivity disorder (ADHD) treatment.
 Suryaprakash Sambhara, DVM, Ph.D
Suryaprakash Sambhara, DVM, Ph.D
Chief, Immunology Section, Influenza Division, CDC, Atlanta, USA
Making sense of pathogen sensors of Innate Immunity: Utility of their ligands as antiviral agents and adjuvants for vaccines.
Currently used antiviral agents act by inhibiting viral entry, replication, or release of viral progeny. However, recent emergence of drug-resistant viruses has become a major public health concern as it is limiting our ability to prevent and treat viral diseases. Furthermore, very few antiviral agents with novel modes of action are currently in development. It is well established that the innate immune system is the first line of defense against invading pathogens. The recognition of diverse pathogen-associated molecular patterns (PAMPs) is accomplished by several classes of pattern recognition receptors (PRRs) and the ligand/receptor interactions trigger an effective innate antiviral response. In the past several years, remarkable progress has been made towards understanding both the structural and functional nature of PAMPs and PRRs. As a result of their indispensable role in virus infection, these ligands have become potential pharmacological agents against viral infections. Since their pathways of action are evolutionarily conserved, the likelihood of viruses developing resistance to PRR activation is diminished. I will discuss the recent developments investigating the potential utility of the ligands of innate immune receptors as antiviral agents and molecular adjuvants for vaccines.

Manjunath Joshi, Apoorva Lad, Bharat Prasad Alevoor, Aswath Balakrishnan, Lingadakai Ramachandra and Kapaettu Satyamoorthy
Pathological conditions during Type 2 Diabetes (T2D) are associated with elevated risk for common community acquired infections due to poor glycemic control. Multiple studies have indicated specific defects in innate and adaptive immune function in diabetic subjects. Neutrophils play an important role in eliminating pathogens as an active constituent of innate immune system. Apart from canonically known phagocytosis mechanism, neutrophils are endowed with a unique ability to produce extracellular traps (NETs) to kill pathogens by expelling DNA coated with bactericidal proteins and histone. NETosis is stimulated by diverse bacteria and their products, fungi, protozoans, cytokines, phorbol esters and by activated platelets. Considering deregulation of metabolic and immune response pathways during pathological state of diabetes and NETosis as a potential mechanism for killing bacteria, we therefore, investigated whether hyperglycemic conditions modulate formation of neutrophil NETs and attempted to identify underlying immunoregulatory mechanisms. Freshly isolated neutrophils from normal individuals were cultured in absence or presence of high glucose (different concentrations) for 24 hours and activated with either LPS (2 mg/ml) or PMA (20 ng/ml) or IL-6 (20 ng/ml) for 3 hours. NETs were visualized and quantified by addition of DNA binding dye SYTOX green using fluorescence microscope and fluorimetry. NETs were quantified in Normal and diabetic subjects. Serum IL-6 levels were measured using ELISA technique. NETs bound elasatse were quantified in normal and diabetic subjects in presence or absence of DNase. Bacterial killing assays were performed upon infecting E.coli with activated neutrophils from normal and diabetic subjects. Microscopy and fluorimetry analysis suggested dramatic impairment in NETs formation under high glucose conditions. Extracellular DNA lattices formed in hyperglycemic conditions were short lived and unstable leading to rapid disintegration. Subsequent, time course experiments showed that NETs production was delayed in hyperglycemic conditions. To validate our findings more closely to clinical conditions, we investigated the neutrophil activation and NETs formation in diabetic patients. Upon stimulation with LPS for three hours, neutrophils from diabetic subjects responded weakly to LPS and lesser NETs were formed; whereas, neutrophils from normal individuals showed robust release of NETs. In few patients we found short and imperfect NETs in basal conditions suggesting constitutive activation of neutrophils in diabetic subjects. Interestingly, NETs bound elastase activity was reduced in diabetes subjects when compared to non-diabetic individuals, indicating a dysfunction of one of the important protein component of NETs during diabetes. Neutrophils from diabetic subjects released higher levels of IL-6 without any stimulation suggesting an existence of constitutively activated pro-inflammatory state. IL-6 induced NETs formation and was abrogated by high glucose. Weobserved that glycolysis inhibitor 2-DG resensitize the high glucose attenuated LPS and IL-6 induced NETs. a) NETs are influenced by glucose homeostasis, b) IL-6 as potent inducer of energy dependent NETs formation and c) hyperglycemia mimics a state of constitutively active pro-inflammatory condition in neutrophils leading to reduced response to external stimuli making diabetic subjects susceptible for infections.







