 S. Ramaswamy, Ph.D.
S. Ramaswamy, Ph.D.
CEO of c-CAMP, Dean, inStem, NCBS, Bangalore, India
Discovery, engineering and applications of Blue Fish Protein with Red Fluorescence
Swagatha Ghosh, Chi-Li Yu, Daniel Ferraro, Sai Sudha, Wayne Schaefer, David T Gibson and S. Ramaswamy
Fluorescent proteins and their applications have revolutionized our understanding of biology significantly. In spite of several years since the discovery of the classic GFP, proteins of this class are used as the standard flag bearers. We have recently discovered a protein from the fish Sanders vitrius that shows interesting fluorescent properties – including a 280 nm stoke shift and infrared emission. The crystal structure of the wild type protein shows that it is a tetramer. We have engineered mutations to make a monomer with very similar fluorescent properties. We have used this protein for tissue imaging as well as for in cell-fluorescence successfully
 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
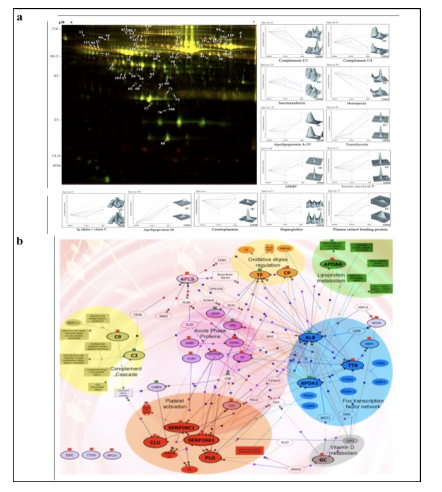
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)
 Nader Pourmand, Ph.D.
Nader Pourmand, Ph.D.
Director, UCSC Genome Technology Center,University of California, Santa Cruz
Biosensor and Single Cell Manipulation using Nanopipettes
Approaching sub-cellular biological problems from an engineering perspective begs for the incorporation of electronic readouts. With their high sensitivity and low invasiveness, nanotechnology-based tools hold great promise for biochemical sensing and single-cell manipulation. During my talk I will discuss the incorporation of electrical measurements into nanopipette technology and present results showing the rapid and reversible response of these subcellular sensors to different analytes such as antigens, ions and carbohydrates. In addition, I will present the development of a single-cell manipulation platform that uses a nanopipette in a scanning ion-conductive microscopy technique. We use this newly developed technology to position the nanopipette with nanoscale precision, and to inject and/or aspirate a minute amount of material to and from individual cells or organelle without comprising cell viability. Furthermore, if time permits, I will show our strategy for a new, single-cell DNA/ RNA sequencing technology that will potentially use nanopipette technology to analyze the minute amount of aspirated cellular material.
 Satheesh Babu T. G., Ph.D.
Satheesh Babu T. G., Ph.D.
Associate Professor, Department of Sciences, School of Engineering, Amrita University, Coimbatore, India
Nanomaterials for ‘enzyme-free’ biosensing
Enzyme based sensors have many draw backs such as poor storage stability, easily affected by the change in pH and temperature and involves complicated enzyme immobilization procedures. To address this limitation, an alternative approach without the use of enzyme, “non-enzymatic” has been tried recently. Choosing the right catalyst for direct electrochemical oxidation / reduction of a target molecule is the key step in the fabrication of non-enzymatic sensors.
Non-enzymatic sensors for glucose, creatinine, vitamins and cholesterol are fabricated using different nanomaterials, such as nanotubes, nanowires and nanoparticles of copper oxide, titanium dioxide, tantalum oxide, platinum, gold and graphenes. These sensors selectively catalyse the targeted analyte with very high sensitivity. These nanomaterials based sensors combat the drawbacks of enzymatic sensors.

H S M Kort, J-W J Lammers, S N W Vorrink, T Troosters
Introduction
Chronic Obstructive Pulmonary Disease (COPD) is a disabling airway disease with variable extrapulmonary effects that may contribute to disease severity in individual patients (Rabe et al. 2007). The world health organization predicts that COPD will become the third leading cause of death worldwide by 2030. Patients with COPD demonstrate reduced levels of spontaneous daily physical activity (DPA) compared with healthy controls (Pitta et al. 2005). This results in a higher risk of hospital admission and shorter survival (Pitta et al. 2006). Pulmonary rehabilitation can help to improve the DPA level, however, obtained benefits decline after 1–2 years (Foglio et al. 2007).
Purpose
In order to maintain DPA in COPD patients after rehabilitation, we developed a mobile phone application. This application measures DPA as steps per day, measured by the accelerometer of the smartphone, and shows the information to the patient via the display of the mobile phone. A physiotherapist can monitor the patient via a secure website where DPA measurements are visible for all patients. Here, DPA goals can be adjusted and text messages sent.
Method
Three pilot studies were performed with healthy students and COPD patients to test the application for usability, user friendliness and reliability with questionnaires and focus groups. Subjects also wore a validated accelerometer. For the Randomized Controlled Trial (RCT) 140 COPD patients will be recruited in Dutch physiotherapy practises. They will be randomised in an intervention group that receives the smartphone for 6 months and a control group. Measurements include lungfunction, dyspnea, and exercise capacity and are held at 0, 3, 6 and 12 months.
Results and Discussion
The application was found to be useful, easy to learn and use. Subjects had no problems with health care professionals seeing information on their physical activity performance. They do find it important to be able to determine who can see the information. Correlations between the accelerometer and the measurements on DPA of the smartphone for steps per hour were 0.69 and 0.70 for pilot studies 1 (students) and 2 (COPD patients) respectively. The version of the application in pilot study 3 contained an error, which made correlations with the accelerometer unusable. The RCT study is now being executed.

Anupama Natarajan, James Hickman and Peter Molnar
Novel Cell-Based Biosensors for High Throughput Toxin Detection and Drug Screening Applications
Over the last decade there has been focus on the development of cellbased biosensors to detect environmental toxins or to combat the threats of biological warfare. These sensors have been shown to have multiple applications including understanding function and behaviour at the cellular and tissue levels, in cell electrophysiology and as drug screening tools that can eliminate animal testing. These factors make the development of cell-based biosensors into high throughput systems a priority in pharmacological, environmental and defence industries (Pancrazio J J et al. 1999, Kang G et al. 2009, Krinke D et al. 2009). We have developed a high through-put in vitro cell-silicon hybrid platform that could be used to analyze both cell function and response to various toxins and drugs. Our hypothesis was that by utilizing surface modification to provide external guidance cues as well as optimal growth conditions for different cell types (Cardiac and Neuronal), we could enhance the information output and content of such a system. An intrinsic part of this study was to create ordered or patterned functional networks of cells on Micro-electrode arrays (MEA). Such engineered networks had a two-fold purpose in that they not only aided in a more accurate analysis of cell response and cell and tissue behaviour, but also increased the efficiency of the system by increasing the connectivity and placement of the cells over the recording electrodes. Here we show the response of this system to various toxins and drugs and the measurement of several vital cardiac parameters like conduction velocity and refractory period (Natarajan A et al. 2011)

Tejaswini Subbannayya, Nandini A. Sahasrabuddhe, Arivusudar Marimuthu, Santosh Renuse, Gajanan Sathe, Srinivas M. Srikanth, Mustafa A. Barbhuiya, Bipin Nair, Juan Carlos Roa, Rafael Guerrero-Preston, H. C. Harsha, David Sidransky, Akhilesh Pandey, T. S. Keshava Prasad and Aditi Chatterjee
Proteomic profiling of gallbladder cancer secretome – a source for circulatory biomarker discovery
Gallbladder cancer (GBC) is the fifth most common cancer of the gastrointestinal tract and one of the common malignancies that occur in the biliary tract (Misra et al. 2006; Lazcano-Ponce et al. 2001). It has a poor prognosis with survival of less than 5 years in 90% of the cases (Misra et al. 2003). The etiology is ill-defined. Several risk factors have been reported including cholelithiasis, obesity, female gender and exposure to carcinogens (Eslick 2010; Kumar et al. 2006). Poor prognosis in GBC is mainly due to late presentation of the disease and lack of reliable biomarkers for early diagnosis. This emphasizes the need to identify and characterize cancer biomarkers to aid in the diagnosis and prognosis of GBC. Secreted proteins are an important class of molecules which can be detected in body fluids and has been targeted for biomarker discovery. There are challenges faced in the proteomic interrogation of body fluids especially plasma such as low abundance of tumor secreted proteins, high complexity and high abundance of other proteins that are not released by the tumor cells (Tonack et al. 2009). Profiling of conditioned media from the cancer cell lines can be used as an alternate means to identify secreted proteins from tumor cells (Kashyap et al. 2010; Marimuthu et al. 2012). We analyzed the invasive property of 7 GBC cell lines (SNU-308, G-415, GB-d1, TGBC2TKB, TGBC24TKB, OCUG-1 and NOZ). Four cell lines were selected for analysis of the cancer secretome based on the invasive property of the cells. We employed isobaric tags for relative and absolute quantitation (iTRAQ) labeling technology coupled with high resolution mass spectrometry to identify and characterize secretome from the panel of 4GBC cancer cells mentioned above. In total, we have identified around 2,000 proteins of which 175 were secreted at differential abundance across all the four cell lines. This secretome analysis will act as a reservoir of candidate biomarkers. Currently, we are investigating and validating these candidate markers from GBC cell secretome. Through this study, we have shown mass spectrometry-based quantitative proteomic analysis as a robust approach to investigate secreted proteins in cancer cells.












