 S. Ramaswamy, Ph.D.
S. Ramaswamy, Ph.D.
CEO of c-CAMP, Dean, inStem, NCBS, Bangalore, India
Discovery, engineering and applications of Blue Fish Protein with Red Fluorescence
Swagatha Ghosh, Chi-Li Yu, Daniel Ferraro, Sai Sudha, Wayne Schaefer, David T Gibson and S. Ramaswamy
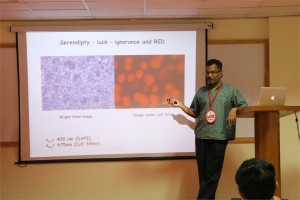
Fluorescent proteins and their applications have revolutionized our understanding of biology significantly. In spite of several years since the discovery of the classic GFP, proteins of this class are used as the standard flag bearers. We have recently discovered a protein from the fish Sanders vitrius that shows interesting fluorescent properties – including a 280 nm stoke shift and infrared emission. The crystal structure of the wild type protein shows that it is a tetramer. We have engineered mutations to make a monomer with very similar fluorescent properties. We have used this protein for tissue imaging as well as for in cell-fluorescence successfully
 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
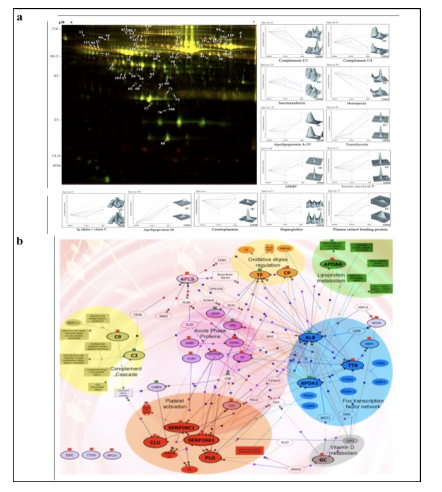
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)
 Shantikumar Nair, Ph.D.
Shantikumar Nair, Ph.D.
Professor & Director, Amrita Center for Nanosciences & Molecular Medicine, Amrita University, India
Spatially Distributed and Hierarchical Nanomaterials in Biotechnology
Although nano materials are well investigated in biotechnology in their zero-, one- and two-dimensional forms, three-dimensional nanomaterials are relatively less investigated for their biological applications. Three dimensional nano materials are much more complex with several structural and hierarchical variables controlling their mechanical, chemical and biological functionality. In this talk examples are given of some complex three dimensional systems including, scaffolds, aggregates, fabrics and membranes. Essentially three types of hierarchies are considered: one-dimensional hierarchy, two-dimensional hierarchy and three-dimensional hierarchy each giving rise to unique behaviors.
 Nader Pourmand, Ph.D.
Nader Pourmand, Ph.D.
Director, UCSC Genome Technology Center,University of California, Santa Cruz
Biosensor and Single Cell Manipulation using Nanopipettes
Approaching sub-cellular biological problems from an engineering perspective begs for the incorporation of electronic readouts. With their high sensitivity and low invasiveness, nanotechnology-based tools hold great promise for biochemical sensing and single-cell manipulation. During my talk I will discuss the incorporation of electrical measurements into nanopipette technology and present results showing the rapid and reversible response of these subcellular sensors to different analytes such as antigens, ions and carbohydrates. In addition, I will present the development of a single-cell manipulation platform that uses a nanopipette in a scanning ion-conductive microscopy technique. We use this newly developed technology to position the nanopipette with nanoscale precision, and to inject and/or aspirate a minute amount of material to and from individual cells or organelle without comprising cell viability. Furthermore, if time permits, I will show our strategy for a new, single-cell DNA/ RNA sequencing technology that will potentially use nanopipette technology to analyze the minute amount of aspirated cellular material.
 Rajgopal Srinivasan, Ph.D.
Rajgopal Srinivasan, Ph.D.
Principal Scientist & Head Bio IT R&D, TCS Innovation Labs, India
Interpretation of Genomic Variation – Identifying Rare Variations Leading to Disease
Genome sequencing technologies are generating an abundance of data on human genetic variations. A big challenge lies in interpreting the functional relevance of such variations, especially in clinical settings. A first step in understanding the clinical relevance of genetic variations is to annotate the variants for region of occurrence, degree of conservation both within and across species, pattern of variation across related individuals, novelty of the variation and know effects of related variations. Several tools already exist for this purpose. However, these tools have their strengths and weaknesses. A second issue is the development of algorithms, which, given a rich annotation of variants are able to prioritize the variants as being relevant to the phenotype under investigation.
In my talk I will detail work that has been done in our labs to address both of the above problems. I will also illustrate the application of these tools that helped identify a rare mutation in the ATM gene leading to a diagnosis of AT in two infants.
 Sharmila Mande, Ph.D.
Sharmila Mande, Ph.D.
Principal Scientist and Head, Bio Sciences R&D, TCS Innovation Labs, Pune
Gut microbiome and health: Moving towards the new era of translational medicine
The microbes inhabiting our body outnumber our own cells by a factor of 10. The genomes of these microbes, called the ‘second genome’ are therefore expected to have great influence on our health and well being. The emerging field of metagenomics is rapidly becoming the method of choice for studying the microbial community (called microbiomes) present in various parts of the human body. Recent studies have implicated the role of gut microbiomes in several diseases and disorders. Studies have indicated gut microbiome’s role in nutrient absorption, immuno-modulation motor-response, and other key physiological processes. However, our understanding of the role of gut microbiota in malnutrition is currently incomplete. Exploration of these aspects are likely to help in understanding the microbial basis for several physiological disorders associated with malnutrition (eg, increased susceptibility to diarrhoeal pathogens) and may finally aid in devising appropriate probiotic strategies addressing this menace. A metagenomic approach was employed for analysing the differences between gut microbial communities obtained from malnourished and healthy children. Results of the analysis using TCS’ ‘Metagenomic Analysis Platform’ were discussed in detail during my talk.
 Lalitha Subramanian, Ph.D.
Lalitha Subramanian, Ph.D.
Chief Scientific Officer & VP, Services at Scienomics, USA
Nanoscale Simulations – Tackling Form and Formulation Challenges in Drug Development and Drug Delivery
Lalitha Subramanian, Dora Spyriouni, Andreas Bick, Sabine Schweizer, and Xenophon Krokidis Scienomics
The discovery of a compound which is potent in activity against a target is a major milestone in Pharmaceutical and Biotech industry. However, a potent compound is only effective as a therapeutic agent when it can be administered such that the optimal quantity is transported to the site of action at an optimal rate. The active pharmaceutical ingredient (API) has to be tested for its physicochemical properties before the appropriate dosage form and formulation can be designed. Some of the commonly evaluated parameters are crystal forms and polymorphs, solubility, dissolution behavior, stability, partition coefficient, water sorption behavior, surface properties, particle size and shape, etc. Pharmaceutical development teams face the challenge of quickly and efficiently determining a number of properties with small quantities of the expensive candidate compounds. Recently the trend has been to screen these properties as early as possible and often the candidate compounds are not available in sufficient quantities. Increasingly, these teams are leveraging nanoscale simulations similar to those employed by drug discovery teams for several decades. Nanoscale simulations are used to predict the behavior using very little experimental data and only if this is promising further experiments are done. Another aspect where nanoscale simulations are being used in drug development and drug delivery is to get insights into the behavior of the system so that process failures can be remediated and formulation performance can be improved. Thus, the predictive screening and the in-depth understanding leads to experimental efficiency resulting in far-reaching business impacts.
With specific examples, this talk will focus on the different types of nanoscale simulations used to predict properties of the API in excipients and also provide insight into system behavior as a function of shelf life, temperature, mechanical stress, etc.
 Satheesh Babu T. G., Ph.D.
Satheesh Babu T. G., Ph.D.
Associate Professor, Department of Sciences, School of Engineering, Amrita University, Coimbatore, India
Nanomaterials for ‘enzyme-free’ biosensing
Enzyme based sensors have many draw backs such as poor storage stability, easily affected by the change in pH and temperature and involves complicated enzyme immobilization procedures. To address this limitation, an alternative approach without the use of enzyme, “non-enzymatic” has been tried recently. Choosing the right catalyst for direct electrochemical oxidation / reduction of a target molecule is the key step in the fabrication of non-enzymatic sensors.
Non-enzymatic sensors for glucose, creatinine, vitamins and cholesterol are fabricated using different nanomaterials, such as nanotubes, nanowires and nanoparticles of copper oxide, titanium dioxide, tantalum oxide, platinum, gold and graphenes. These sensors selectively catalyse the targeted analyte with very high sensitivity. These nanomaterials based sensors combat the drawbacks of enzymatic sensors.
 Srisairam Achuthan, Ph.D.
Srisairam Achuthan, Ph.D.
Senior Scientific Programmer, Research Informatics Division, Department of Information Sciences, City of Hope, CA, USA
Applying Machine learning for Automated Identification of Patient Cohorts
Srisairam Achuthan, Mike Chang, Ajay Shah, Joyce Niland
Patient cohorts for a clinical study are typically identified based on specific selection criteria. In most cases considerable time and effort are spent in finding the most relevant criteria that could potentially lead to a successful study. For complex diseases, this process can be more difficult and error prone since relevant features may not be easily identifiable. Additionally, the information captured in clinical notes is in non-coded text format. Our goal is to discover patterns within the coded and non-coded fields and thereby reveal complex relationships between clinical characteristics across different patients that would be difficult to accomplish manually. Towards this, we have applied machine learning techniques such as artificial neural networks and decision trees to determine patients sharing similar characteristics from available medical records. For this proof of concept study, we used coded and non-coded (i.e., clinical notes) patient data from a clinical database. Coded clinical information such as diagnoses, labs, medications and demographics recorded within the database were pooled together with non-coded information from clinical notes including, smoking status, life style (active / inactive) status derived from clinical notes. The non-coded textual information was identified and interpreted using a Natural Language Processing (NLP) tool I2E from Linguamatics.








