Vural Özdemir Ph.D.
Sanjeeva Srivastava Ph.D.
 Upinder S. Bhalla, Ph.D.
Upinder S. Bhalla, Ph.D.
Professor & Dean, NCBS, Bengaluru, India
Watching the network change during the formation of associative memory
The process of learning is measured through behavioural changes, but it is of enormous interest to understand its cellular and network basis. We used 2-photon imaging of hippocampal CA1 pyramidal neuron activity in mice to monitor such changes during the acquisition of a trace conditioning task. One of the questions in such learning is how the network retains a trace of a brief conditioned stimulus (a sound), until the arrival of a delayed unconditioned stimulus (a puff of air to the eye). During learning, the mice learn to blink when the tone is presented, well before the arrival of the air puff.
The mice learnt this task in 20-50 trials. We observed that in this time-frame the cells in the network changed the time of their peak activity, such that their firing times tiled the interval between sound and air puff. Thus the cells seem to form a relay of activity. We also observed an evolution in functional connectivity in the network, as measured by groupings of correlated cells. These groupings were stable till the learning protocol commenced, and then changed. Thus we have been able to observe two aspects of network learning: changes in activity (relay firing), and changes in connectivity (correlation groups).
 Claudia AM Wheeler-Kingshott, Ph.D.
Claudia AM Wheeler-Kingshott, Ph.D.
University Reader in Magnetic Resonance Physics, Department of Neuroinflammation, UCL Institute of Neurology, London, UK
Abstract
Detecting neuronal activity in vivo non-invasively is possible with a number of techniques. Amongst these, in 1990 functional magnetic resonance imaging (fMRI) was proposed as a technique that has a great ability to spatially map brain activity by exploiting the blood oxygenation level dependent (BOLD) contrast mechanism [1, 2]. In fact, neuronal activation triggers a demand for oxygen and induces a localised increase in blood flow and blood volume, which actually exceeds the metabolic needs. This in turns causes an increase of oxyhaemoglobin in the venous compartment, which is a transient phenomenon and is accompanied by a transient change (decrease) in the concentration of deoxyhaemoglobin. Due to its paramagnetic properties, the amount of deoxyhaemoglobin present in the venous blood affects the local magnetic field seen by the spins (protons) and determines the local properties of the MR signal. A decrease in deoxyhaemoglobin during neuronal activity, therefore, induces local variations of this magnetic field that increases the average transverse relaxation time of tissue, measured via the T2* parameter [3]. This means that there is an increase of the MR signal (of the order of a few %, typically <5%) linked to metabolic changes happening during brain function. Activation can be inferred at different brain locations by performing tasks while acquiring the MR signal and comparing periods of rest to periods of activity.
The macroscopic changes of the BOLD signal are well characterised, while the reason for the increased blood supply, exceeding demands, needs further thoughts. Here we will discuss two approaches for explaining the BOLD phenomenon, one that links it to adenosine triphosphate production [4] and enzyme saturation, the other that relates it to the very slow diffusion of oxygen through the blood-brain-barrier with a consequent compensatory high demand of oxygen [5]. Some evidence of restricted oxygen diffusion has been shown by means of hypercapnia [6], although it is not excluded that both mechanisms may be present.
Overall, the BOLD signal changes theory and its physiological basis will be presented and discussed.
References
- Ogawa, S., et al., Brain magnetic resonance imaging with contrast dependent on blood oxygenation. Proc Natl Acad Sci U S A, 1990. 87(24): p. 9868-72.
- Kwong, K.K., et al., Dynamic magnetic resonance imaging of human brain activity during primary sensory stimulation. Proc Natl Acad Sci U S A, 1992. 89(12): p. 5675-9.
- Bandettini PA, et al. Spin-echo and gradient-echo EPI of human brain activation using BOLD contrast: a comparative study at 1.5 T. NMR Biomed. 1994 Mar;7(1-2):12-20
- Fox, P.T., et al., Nonoxidative glucose consumption during focal physiologic neural activity. Science, 1988. 241(4864): p. 462-4.
- Gjedde, A., et al. Reduction of functional capillary density in human brain after stroke. J Cereb Blood Flow Metab, 1990. 10(3): p. 317-26.
- Hoge, R.D., et al., Linear coupling between cerebral blood flow and oxygen consumption in activated human cortex. Proc Natl Acad Sci U S A, 1999. 96(16): p. 9403-8.
 Aditya Murthy, Ph.D.
Aditya Murthy, Ph.D.
Associate Professor, Centre For Neuroscience, Indian Institute of Science, Bangalore, India
Since Karl Lashley’s seminal work on the formulation of serial order, numerous models assume simultaneous representation of competitive elements of a sequence, to account for serial order effects in different types of behavior like typing, speech, etc. Such models follow two basic assumptions: (1) more than one plan representation can be simultaneously active in a planning layer; (2) the most active plan is chosen in another layer called the competitive choice layer. Using the oculomotor system I will describe behavioral and neurophysiological experiments that tests the two critical predictions of such queuing models, providing evidence that basal ganglia in monkeys and humans instantiate a form of queuing that transforms parallel movement representations into more serial representations, allowing for the expression of sequential saccadic eye movements.
 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
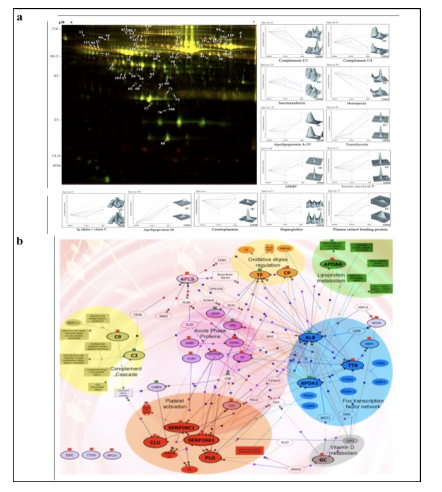
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)
 Nader Pourmand, Ph.D.
Nader Pourmand, Ph.D.
Director, UCSC Genome Technology Center,University of California, Santa Cruz
Biosensor and Single Cell Manipulation using Nanopipettes
Approaching sub-cellular biological problems from an engineering perspective begs for the incorporation of electronic readouts. With their high sensitivity and low invasiveness, nanotechnology-based tools hold great promise for biochemical sensing and single-cell manipulation. During my talk I will discuss the incorporation of electrical measurements into nanopipette technology and present results showing the rapid and reversible response of these subcellular sensors to different analytes such as antigens, ions and carbohydrates. In addition, I will present the development of a single-cell manipulation platform that uses a nanopipette in a scanning ion-conductive microscopy technique. We use this newly developed technology to position the nanopipette with nanoscale precision, and to inject and/or aspirate a minute amount of material to and from individual cells or organelle without comprising cell viability. Furthermore, if time permits, I will show our strategy for a new, single-cell DNA/ RNA sequencing technology that will potentially use nanopipette technology to analyze the minute amount of aspirated cellular material.
 Vural Özdemir, MD, Ph.D., DABCP
Vural Özdemir, MD, Ph.D., DABCP
Co-Founder, DELSA Global, Seattle, WA, USA
Crowd-Funded Micro-Grants to Link Biotechnology and “Big Data” R&D to Life Sciences Innovation in India
Vural Özdemir, MD, PhD, DABCP1,2*
- Data-Enabled Life Sciences Alliance International (DELSA Global), Seattle, WA 98101, USA;
- Faculty of Management and Medicine, McGill University, Canada;
ABSTRACT
Aims: This presentation proposes two innovative funding solutions for linking biotechnology and “Big Data” R&D in India with artisan small scale discovery science, and ultimately, with knowledge-based innovation:
- crowd-funded micro-grants, and
- citizen philanthropy
These two concepts are new, and inter-related, and can be game changing to achieve the vision of biotechnology innovation in India, and help bridge local innovation with global science.
Background and Context: Biomedical science in the 21(st) century is embedded in, and draws from, a digital commons and “Big Data” created by high-throughput Omics technologies such as genomics. Classic Edisonian metaphors of science and scientists (i.e., “the lone genius” or other narrow definitions of expertise) are ill equipped to harness the vast promises of the 21(st) century digital commons. Moreover, in medicine and life sciences, experts often under-appreciate the important contributions made by citizen scholars and lead users of innovations to design innovative products and co-create new knowledge. We believe there are a large number of users waiting to be mobilized so as to engage with Big Data as citizen scientists-only if some funding were available. Yet many of these scholars may not meet the meta-criteria used to judge expertise, such as a track record in obtaining large research grants or a traditional academic curriculum vitae. This presentation will describe a novel idea and action framework: micro-grants, each worth $1000, for genomics and Big Data. Though a relatively small amount at first glance, this far exceeds the annual income of the “bottom one billion” – the 1.4 billion people living below the extreme poverty level defined by the World Bank ($1.25/day).
We will present two types of micro-grants. Type 1 micro-grants can be awarded through established funding agencies and philanthropies that create micro-granting programs to fund a broad and highly diverse array of small artisan labs and citizen scholars to connect genomics and Big Data with new models of discovery such as open user innovation. Type 2 micro-grants can be funded by existing or new science observatories and citizen think tanks through crowd-funding mechanisms described herein. Type 2 micro-grants would also facilitate global health diplomacy by co-creating crowd-funded micro-granting programs across nation-states in regions facing political and financial instability, while sharing similar disease burdens, therapeutics, and diagnostic needs. We report the creation of ten Type 2 micro-grants for citizen science and artisan labs to be administered by the nonprofit Data-Enabled Life Sciences Alliance International (DELSA Global, Seattle: http://www.delsaglobal.org). Our hope is that these micro-grants will spur novel forms of disruptive innovation and life sciences translation by artisan scientists and citizen scholars alike.
Address Correspondence to:
Vural Özdemir, MD, PhD, DABCP
Senior Scholar and Associate Professor
Faculty of Management and Medicine, McGill University
1001 Sherbrooke Street West
Montreal, Canada H3A 1G5








