 K. Satyamoorthy, Ph.D.
K. Satyamoorthy, Ph.D.
Director, Life Sciences Centre, Manipal University, India
Epigenetic Changes due to DNA Methylation in Human Epithelial Tumors
Extensive global hypomethylation in the genome and hypermthylation of selective tumor specific suppressor genes appears to be a hallmark of human cancers. Data suggests that hypermethylation of promoter region in genes is more closely related to subsequent gene expression; contrary to gene-body DNA methylation. The intricate balance between these two may contribute to the progressive process of development, differentiation and carcinogenesis. Epigenetic changes encompass, apart from DNA methylation, chromatin modifications through post-translational changes in histones and control by miRNAs. At the genome level, effects from these are compounded by copy number variations (CNVs) which may ultimately influence protein functions. From clinical perspective, changes in DNA methylation occur very early which are reversible and are influenced by environmental factors. Therefore, these can be potential resource for identifying therapeutic targets as well as biomarkers for early screening of cancer. Our current efforts in profiling genome wide DNA methylation changes in oral, cervical and breast cancers through DNA methylation microarray analysis has revealed number of alterations critical for survival, progression and metastatic behavior of tumors. Bioinformatics and functional analysis revealed several key regulatory molecules controlled by DNA methylation and suggests that DNA methylation changes in several CpG islands appear to co-segregate in the regions of miRNAs as well as in the CNVs. We have validated the signatures for methylation of CpG islands through bisufite sequencing for essential genes in clinical samples and have undertaken transcriptional and functional analysis in tumor cell lines. These results will be presented.
 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
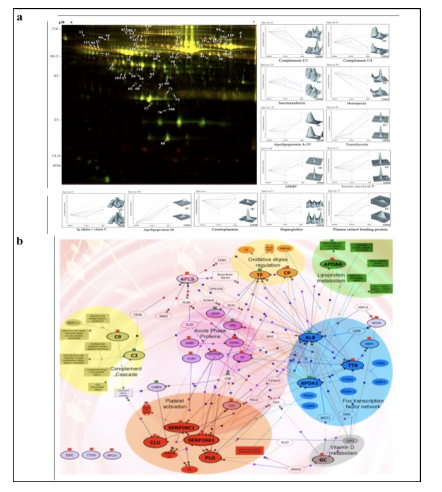
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)
 Nader Pourmand, Ph.D.
Nader Pourmand, Ph.D.
Director, UCSC Genome Technology Center,University of California, Santa Cruz
Biosensor and Single Cell Manipulation using Nanopipettes
Approaching sub-cellular biological problems from an engineering perspective begs for the incorporation of electronic readouts. With their high sensitivity and low invasiveness, nanotechnology-based tools hold great promise for biochemical sensing and single-cell manipulation. During my talk I will discuss the incorporation of electrical measurements into nanopipette technology and present results showing the rapid and reversible response of these subcellular sensors to different analytes such as antigens, ions and carbohydrates. In addition, I will present the development of a single-cell manipulation platform that uses a nanopipette in a scanning ion-conductive microscopy technique. We use this newly developed technology to position the nanopipette with nanoscale precision, and to inject and/or aspirate a minute amount of material to and from individual cells or organelle without comprising cell viability. Furthermore, if time permits, I will show our strategy for a new, single-cell DNA/ RNA sequencing technology that will potentially use nanopipette technology to analyze the minute amount of aspirated cellular material.
 Rajgopal Srinivasan, Ph.D.
Rajgopal Srinivasan, Ph.D.
Principal Scientist & Head Bio IT R&D, TCS Innovation Labs, India
Interpretation of Genomic Variation – Identifying Rare Variations Leading to Disease
Genome sequencing technologies are generating an abundance of data on human genetic variations. A big challenge lies in interpreting the functional relevance of such variations, especially in clinical settings. A first step in understanding the clinical relevance of genetic variations is to annotate the variants for region of occurrence, degree of conservation both within and across species, pattern of variation across related individuals, novelty of the variation and know effects of related variations. Several tools already exist for this purpose. However, these tools have their strengths and weaknesses. A second issue is the development of algorithms, which, given a rich annotation of variants are able to prioritize the variants as being relevant to the phenotype under investigation.
In my talk I will detail work that has been done in our labs to address both of the above problems. I will also illustrate the application of these tools that helped identify a rare mutation in the ATM gene leading to a diagnosis of AT in two infants.
 Satheesh Babu T. G., Ph.D.
Satheesh Babu T. G., Ph.D.
Associate Professor, Department of Sciences, School of Engineering, Amrita University, Coimbatore, India
Nanomaterials for ‘enzyme-free’ biosensing
Enzyme based sensors have many draw backs such as poor storage stability, easily affected by the change in pH and temperature and involves complicated enzyme immobilization procedures. To address this limitation, an alternative approach without the use of enzyme, “non-enzymatic” has been tried recently. Choosing the right catalyst for direct electrochemical oxidation / reduction of a target molecule is the key step in the fabrication of non-enzymatic sensors.
Non-enzymatic sensors for glucose, creatinine, vitamins and cholesterol are fabricated using different nanomaterials, such as nanotubes, nanowires and nanoparticles of copper oxide, titanium dioxide, tantalum oxide, platinum, gold and graphenes. These sensors selectively catalyse the targeted analyte with very high sensitivity. These nanomaterials based sensors combat the drawbacks of enzymatic sensors.

Pandiaraj Manickam, Niroj Kumar Sethy, Kalpana Bhargava, Vepa Kameswararao and Karunakaran Chandran
Designing electrochemical label free immunosensors for cytochrome c using nanocomposites functionalized screen printed electrodes
Release of cytochrome c (cyt c) from mitochondria into cytosol is a hallmark of apoptosis, used as a biomarker of mitochondrial dependent pathway of cell death (Kluck et al. 1997; Green et al. 1998). We have previously reported cytochrome c reductase (CcR) based biosensors for the measurement of mitochondrial cyt c release (Pandiaraj et al. 2013). Here, we describe the development of novel label-free, immunosensor for cyt c utilizing its specific monoclonal antibody. Two types of nanocomposite modified immunosensing platforms were used for the immobilization of anti-cyt c; (i) Self-assembled monolayer (SAM) functionalized gold nanoparticles (GNP) in conducting polypyrrole (PPy) modified screen printed electrodes (SPE) (ii) Carbon nanotubes (CNT) incorporated PPy on SPE. The nanotopologies of the modified electrodes were confirmed by scanning electron microscopy (SEM). Cyclic voltammetry, electrochemical impedance spectroscopy (EIS) were used for probing the electrochemical properties of the nanocomposite modified electrodes. Method for cyt c quantification is based on the direct electron transfer between Fe3+/Fe2+-heme of cyt c selectively bound to anti-cyt c modified electrode. The Faradaic current response of these nanoimmunosensor increases with increase in cyt c concentration. The procedure for cyt c detection was also optimized (pH, incubation times, and characteristics of electrodes) to improve the analytical characteristics of immunosensors. The analytical performance of anti-cyt c biofunctionalized GNP-PPy nanocomposite platform (detection limit 0.5 nM; linear range: 0.5 nM–2 μM) was better than the CNT-PPy (detection limit 2 nM; linear range: 2 nM-500nM). The detection limits were well below the normal physiological concentration range (Karunakaran et al. 2008). The proposed method does not require any signal amplification or labeled secondary antibodies contrast to widespread ELISA and Western blot. The immunosensors results in simple and rapid measurement of cyt c and has great potential to become an inexpensive and portable device for conventional clinical immunoassays.

Anupama Natarajan, James Hickman and Peter Molnar
Novel Cell-Based Biosensors for High Throughput Toxin Detection and Drug Screening Applications
Over the last decade there has been focus on the development of cellbased biosensors to detect environmental toxins or to combat the threats of biological warfare. These sensors have been shown to have multiple applications including understanding function and behaviour at the cellular and tissue levels, in cell electrophysiology and as drug screening tools that can eliminate animal testing. These factors make the development of cell-based biosensors into high throughput systems a priority in pharmacological, environmental and defence industries (Pancrazio J J et al. 1999, Kang G et al. 2009, Krinke D et al. 2009). We have developed a high through-put in vitro cell-silicon hybrid platform that could be used to analyze both cell function and response to various toxins and drugs. Our hypothesis was that by utilizing surface modification to provide external guidance cues as well as optimal growth conditions for different cell types (Cardiac and Neuronal), we could enhance the information output and content of such a system. An intrinsic part of this study was to create ordered or patterned functional networks of cells on Micro-electrode arrays (MEA). Such engineered networks had a two-fold purpose in that they not only aided in a more accurate analysis of cell response and cell and tissue behaviour, but also increased the efficiency of the system by increasing the connectivity and placement of the cells over the recording electrodes. Here we show the response of this system to various toxins and drugs and the measurement of several vital cardiac parameters like conduction velocity and refractory period (Natarajan A et al. 2011)

Manjunath Joshi, Apoorva Lad, Bharat Prasad Alevoor, Aswath Balakrishnan, Lingadakai Ramachandra and Kapaettu Satyamoorthy
Pathological conditions during Type 2 Diabetes (T2D) are associated with elevated risk for common community acquired infections due to poor glycemic control. Multiple studies have indicated specific defects in innate and adaptive immune function in diabetic subjects. Neutrophils play an important role in eliminating pathogens as an active constituent of innate immune system. Apart from canonically known phagocytosis mechanism, neutrophils are endowed with a unique ability to produce extracellular traps (NETs) to kill pathogens by expelling DNA coated with bactericidal proteins and histone. NETosis is stimulated by diverse bacteria and their products, fungi, protozoans, cytokines, phorbol esters and by activated platelets. Considering deregulation of metabolic and immune response pathways during pathological state of diabetes and NETosis as a potential mechanism for killing bacteria, we therefore, investigated whether hyperglycemic conditions modulate formation of neutrophil NETs and attempted to identify underlying immunoregulatory mechanisms. Freshly isolated neutrophils from normal individuals were cultured in absence or presence of high glucose (different concentrations) for 24 hours and activated with either LPS (2 mg/ml) or PMA (20 ng/ml) or IL-6 (20 ng/ml) for 3 hours. NETs were visualized and quantified by addition of DNA binding dye SYTOX green using fluorescence microscope and fluorimetry. NETs were quantified in Normal and diabetic subjects. Serum IL-6 levels were measured using ELISA technique. NETs bound elasatse were quantified in normal and diabetic subjects in presence or absence of DNase. Bacterial killing assays were performed upon infecting E.coli with activated neutrophils from normal and diabetic subjects. Microscopy and fluorimetry analysis suggested dramatic impairment in NETs formation under high glucose conditions. Extracellular DNA lattices formed in hyperglycemic conditions were short lived and unstable leading to rapid disintegration. Subsequent, time course experiments showed that NETs production was delayed in hyperglycemic conditions. To validate our findings more closely to clinical conditions, we investigated the neutrophil activation and NETs formation in diabetic patients. Upon stimulation with LPS for three hours, neutrophils from diabetic subjects responded weakly to LPS and lesser NETs were formed; whereas, neutrophils from normal individuals showed robust release of NETs. In few patients we found short and imperfect NETs in basal conditions suggesting constitutive activation of neutrophils in diabetic subjects. Interestingly, NETs bound elastase activity was reduced in diabetes subjects when compared to non-diabetic individuals, indicating a dysfunction of one of the important protein component of NETs during diabetes. Neutrophils from diabetic subjects released higher levels of IL-6 without any stimulation suggesting an existence of constitutively activated pro-inflammatory state. IL-6 induced NETs formation and was abrogated by high glucose. Weobserved that glycolysis inhibitor 2-DG resensitize the high glucose attenuated LPS and IL-6 induced NETs. a) NETs are influenced by glucose homeostasis, b) IL-6 as potent inducer of energy dependent NETs formation and c) hyperglycemia mimics a state of constitutively active pro-inflammatory condition in neutrophils leading to reduced response to external stimuli making diabetic subjects susceptible for infections.
 Jeff Perry, Ph.D.
Jeff Perry, Ph.D.
Assistant Professor, University of California, Riverside
Combined Crystallography and SAXS Methods for Studying Macromolecular Complexes
Recent developments in small angle X-ray scattering (SAXS) are rapidly providing new insights into protein interactions, complexes and conformational states in solution, allowing for detailed biophysical quantification of samples of interest1. Initial analyses provide a judgment of sample quality, revealing the potential presence of aggregation, the overall extent of folding or disorder, the radius of gyration, maximum particle dimensions and oligomerization state. Structural characterizations may include ab initio approaches from SAXS data alone, or enhance structural solutions when combined with previously determined crystal/NMR domains. This combination can provide definitions of architectures, spatial organizations of the protein domains within a complex, including those not yet determined by crystallography or NMR, as well as defining key conformational states. Advantageously, SAXS is not generally constrained by macromolecule size, and rapid collection of data in a 96-well plate format provides methods to screen sample conditions. Such screens include co-factors, substrates, differing protein or nucleotide partners or small molecule inhibitors, to more fully characterize the variations within assembly states and key conformational changes. These analyses are also useful for screening constructs and conditions that are most likely to promote crystal growth. Moreover, these high throughput structural determinations can be leveraged to define how polymorphisms affect assembly formations and activities. Also, SAXS-based technologies may be potentially used for novel structure-based screening, for compounds inducing shape changes or associations/diassociations. This is addition to defining architectural characterizations of complexes and interactions for systems biology-based research, and distinctions in assemblies and interactions in comparative genomics. Thus, SAXS combined with crystallography/NMR and computation provides a unique set of tools that should be considered as being part of one’s repertoire of biophysical analyses, when conducting characterizations of protein and other macromolecular interactions.
1 Perry JJ & Tainer JA. Developing advanced X-ray scattering methods combined with crystallography and computation. Methods. 2013 Mar;59(3):363-71.




