 Deepthy Menon, Ph.D.
Deepthy Menon, Ph.D.
Associate Professor, Centre for Nanosciences & Molecular Medicine, Health Sciences Campus, Amrita University, Kochi, India
Nanobioengineering of implant materials for improved cellular response and activity
Deepthy Menon, Divyarani V V, Chandini C Mohan, Manitha B Nair, Krishnaprasad C & Shantikumar V Nair
Abstract
Current trends in biomaterials research and development include the use of surfaces with topographical features at the nanoscale (dimensions < 100 nm), which influence biomolecular or cellular level reactions in vitro and in vivo. Progress in nanotechnology now makes it possible to precisely design and modulate the surface properties of materials used for various applications in medicine at the nanoscale. Nanoengineered surfaces, owing to their close resemblance with extracellular matrix, possess the unique capacity to directly affect protein adsorption that ultimately modulates the cellular adhesion and proliferation at the site of implantation. Taking advantage of this exceptional ability, we have nanoengineered metallic surfaces of Titanium (Ti) and its alloys (Nitinol -NiTi), as well as Stainless Steel (SS) by a simple hydrothermal method for generating non-periodic, homogeneous nanostructures. The bio- and hemocompatibility of these nanotextured metallic surfaces suggest their potential use for orthopedic, dental or vascular implants. The applicability of nanotextured Ti implants for orthopedic use was demonstrated in vivo in rat models, wherein early-stage bone formation at the tissue-implant interface without any fibrous tissue intervention was achieved. This nanoscale topography also was found to critically influence bacterial adhesion in vitro, with decreased adherence of staphylococcus aureus. The same surface nanotopography also was found to provide enhanced proliferation and functionality of vascular endothelial cells, suggesting its prospective use for developing an antithrombotic stent surface for coronary applications. Clinical SS & NiTi stents were also modified based on this strategy, which would offer a suitable solution to reduce the probability of late stent thrombosis associated with bare metallic stents. Thus, we demonstrate that nanotopography on implant surfaces has a critical influence on the fate of cells, which in turn dictates the long term success of the implant.
Acknowledgement: Authors gratefully acknowledge the financial support from Department of Biotechnology, Government of India through the Bioengineering program.
 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
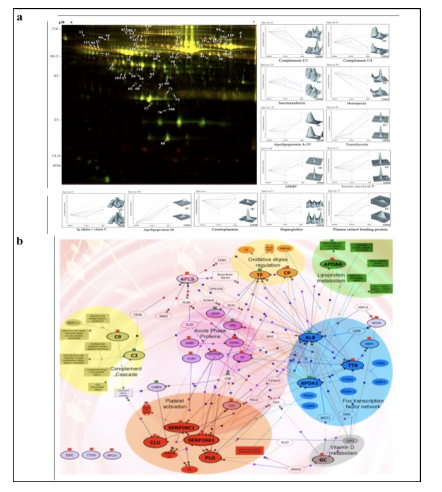
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)
 Nader Pourmand, Ph.D.
Nader Pourmand, Ph.D.
Director, UCSC Genome Technology Center,University of California, Santa Cruz
Biosensor and Single Cell Manipulation using Nanopipettes
Approaching sub-cellular biological problems from an engineering perspective begs for the incorporation of electronic readouts. With their high sensitivity and low invasiveness, nanotechnology-based tools hold great promise for biochemical sensing and single-cell manipulation. During my talk I will discuss the incorporation of electrical measurements into nanopipette technology and present results showing the rapid and reversible response of these subcellular sensors to different analytes such as antigens, ions and carbohydrates. In addition, I will present the development of a single-cell manipulation platform that uses a nanopipette in a scanning ion-conductive microscopy technique. We use this newly developed technology to position the nanopipette with nanoscale precision, and to inject and/or aspirate a minute amount of material to and from individual cells or organelle without comprising cell viability. Furthermore, if time permits, I will show our strategy for a new, single-cell DNA/ RNA sequencing technology that will potentially use nanopipette technology to analyze the minute amount of aspirated cellular material.
 Satheesh Babu T. G., Ph.D.
Satheesh Babu T. G., Ph.D.
Associate Professor, Department of Sciences, School of Engineering, Amrita University, Coimbatore, India
Nanomaterials for ‘enzyme-free’ biosensing
Enzyme based sensors have many draw backs such as poor storage stability, easily affected by the change in pH and temperature and involves complicated enzyme immobilization procedures. To address this limitation, an alternative approach without the use of enzyme, “non-enzymatic” has been tried recently. Choosing the right catalyst for direct electrochemical oxidation / reduction of a target molecule is the key step in the fabrication of non-enzymatic sensors.
Non-enzymatic sensors for glucose, creatinine, vitamins and cholesterol are fabricated using different nanomaterials, such as nanotubes, nanowires and nanoparticles of copper oxide, titanium dioxide, tantalum oxide, platinum, gold and graphenes. These sensors selectively catalyse the targeted analyte with very high sensitivity. These nanomaterials based sensors combat the drawbacks of enzymatic sensors.

Anupama Natarajan, James Hickman and Peter Molnar
Novel Cell-Based Biosensors for High Throughput Toxin Detection and Drug Screening Applications
Over the last decade there has been focus on the development of cellbased biosensors to detect environmental toxins or to combat the threats of biological warfare. These sensors have been shown to have multiple applications including understanding function and behaviour at the cellular and tissue levels, in cell electrophysiology and as drug screening tools that can eliminate animal testing. These factors make the development of cell-based biosensors into high throughput systems a priority in pharmacological, environmental and defence industries (Pancrazio J J et al. 1999, Kang G et al. 2009, Krinke D et al. 2009). We have developed a high through-put in vitro cell-silicon hybrid platform that could be used to analyze both cell function and response to various toxins and drugs. Our hypothesis was that by utilizing surface modification to provide external guidance cues as well as optimal growth conditions for different cell types (Cardiac and Neuronal), we could enhance the information output and content of such a system. An intrinsic part of this study was to create ordered or patterned functional networks of cells on Micro-electrode arrays (MEA). Such engineered networks had a two-fold purpose in that they not only aided in a more accurate analysis of cell response and cell and tissue behaviour, but also increased the efficiency of the system by increasing the connectivity and placement of the cells over the recording electrodes. Here we show the response of this system to various toxins and drugs and the measurement of several vital cardiac parameters like conduction velocity and refractory period (Natarajan A et al. 2011)
 Bodo Eickhoff, Ph.D.
Bodo Eickhoff, Ph.D.
Senior Vice-President, Head of Sales and Marketing for Roche Applied Science, Germany
New paths for treatment of complex diseases: target combinatorial drug therapy
Several types of diseases show a complex pathogenesis and require targeted as well as combinatorial drug treatment. A classical example, Tuberculosis, was thought for decades to be managable by triple therapy, however now requiring new therapeutic approaches due to multi drug resistant strains. HIV and AIDS can only be kept under control by combinations of specific, virus-protein targeted drugs, requiring constant monitoring of resistance patterns and modulation of drug combinations during life-long therapy. As a third example, Cancer in all its different variations, requires detailled molecular understanding to enable targeted therapy. New technologies provide more and in depths molecular insights into pathomechanisms and resulting treatment options. However, is there an alternative way to approach complex diseases by holistic models? Can restoring of apoptosis-capabilities of transformed cells be an example of such an alternative path? How do we in future adress major unresolved topics like increasing drug resistance in bacterial infections, lack of anti-viral drugs, treatment of parasite diseases like Malaria, and newly emerging infectious diseases in research and fast translation of these results into diagnosis and treatment?

Aswath Balakrishnan, Kapaettu Satyamoorthy and Manjunath B Joshi
Introduction
Insulin resistance is a hall mark of metabolic disorders such as diabetes. Reduced insulin response in vasculature leads to disruption of IR/Akt/eNOS signaling pathway resulting in vasoconstriction and subsequently to cardiovascular diseases. Recent studies have demonstrated that inflammatory regulator interleukin-6 (IL-6), as one of the potential mediators that can link chronic inflammation with insulin resistance. Accumulating evidences suggest a significant role of epigenetic mechanisms such as DNA methylation in progression of metabolic disorders. Hence the present study aimed to understand the role of epigenetic mechanisms involved during IL-6 induced vascular insulin resistance and its consequences in cardiovascular diseases.
Materials and Methods
Human umbilical vein endothelial cells (HUVEC) and Human dermal microvascular endothelial cells (HDMEC) were used for this study. Endothelial cells were treated in presence or absence of IL-6 (20ng/ml) for 36 hours and followed by insulin (100nM) stimulation for 15 minutes. Using immunoblotting, cell lysates were stained for phosphor- and total Akt levels to measure insulin resistance. To investigate changes in DNA methylation, cells were treated with or without neutrophil conditioned medium (NCM) as a physiological source of inflammation or IL-6 (at various concentrations) for 36 hours. Genomic DNA was processed for HPLC analysis for methyl cytosine content and cell lysates were analyzed for DNMT1 (DNA (cytosine-5)-methyltransferase 1) and DNMT3A (DNA (cytosine-5)-methyltransferase 3A) levels using immunoblotting.
Results
Endothelial cells stimulated with insulin exhibited an increase in phosphorylation of Aktser 473 in serum free conditions but such insulin response was not observed in cells treated with IL-6, suggesting chronic exposure of endothelial cells to IL-6 leads to insulin resistance. HPLC analysis for global DNA methylation resulted in decreased levels of 5-methyl cytosine in cells treated with pro-inflammatory molecules (both by NCM and IL-6) as compared to untreated controls. Subsequently, analysis in cells treated with IL-6 showed a significant decrease in DNMT1 levels but not in DNMT3A. Other pro-inflammatory marker such as TNF-α did not exhibit such changes.
Conclusion
Our study suggests: a) Chronic treatment of endothelial cells with IL-6 results in insulin resistance b) Neutrophil conditioned medium and IL-6 decreases methyl cytosine levels c) DNMT1 but not DNMT3a levels are reduced in cells treated with IL-6.






