 Ayyappan Nair, Ph.D.
Ayyappan Nair, Ph.D.
Head, Business Development (Technologies, Discovery Biology), Anthem Biosciences & DavosPharma, New Jersey, USA
Inhibition of NF-κB regulated gene expression by chrysoeriol suppresses tumorigenesis in breast cancer cells
Amrutha K1, Pandurangan Nanjan1, Sanu K Shaji1, Damu Sunilkumar1, Subhalakshmi K1, Rashmi U Nair1, Lakshmi Rajakrishna2, Asoke Banerji1, Ayyappan Ramesh Nair1*,2
- School of Biotechnology, Amrita Vishwa Vidyapeetham, Amritapuri Campus, Clappana P.O., Kollam – 690 525, Kerala, India
- Anthem Biosciences, No 49, Canara Bank Road, Bommasandra Industrial Area, Phase 1, Hosur Road, Bangalore – 560 099, Karnataka, India
Abstract: A large number of effective cancer-preventing compounds inhibit the activation of nuclear factor-κ B (NF-κB). It has been previously demonstrated that some flavonoids that are a vital component of our diet inhibits this pathway. As a consequence, many flavonoids inhibit genes involved in various aspects of tumorigenesis and have thus emerged as potential chemopreventive candidates for cancer treatment. We studied the effect of 17 different flavonoids, including the highly evaluated quercetin on the NF-κB pathway, and on the expression of MMP-9 and COX-2 (two NF-κB regulated genes involved in metastasis) in the highly invasive human breast cancer cell line MDA-MB-231. The findings suggest that not all the quercetin like flavone backbone compounds inhibit the NF-κB pathway, and that the highly hydoxylated flavonols quercetagetin and gossypetin did not inhibit this pathway, nor did it inhibit the expression of MMP-9 and COX-2. This indicates a correlation between inhibition of NF-κB and subsequent suppression of these NF-κB regulated genes. Here, we also report the novel observation that the not so well characterized methoxylated flavone chrysoeriol inhibited the NF-κB pathway, and was most potent in reducing the expression of MMP-9 and COX-2. Based on these observations, the cellular effects of chrysoeriol were evaluated in MDA-MB-231. Chrysoeriol caused cell cycle arrest at G2/M, inhibited migration and invasion, and caused cell death of macrophages that contributed to migration of these cancer cells. These effects of chrysoeriol make it a potential therapeutic candidate for breast cancer metastasis.
 Pradip K. Bhatnagar, Ph.D.
Pradip K. Bhatnagar, Ph.D.
Former President & Head, Daiichi Sankyo Life Science Research Centre, India
Strategies for Diseases/Target Selection for Drug Discovery and a Multi-Targeted Approach to Metabolic Disorder
Drug discovery and development is a high risk and expensive undertaking. Although, technologies, such as, bioinformatics, genomics, high throughput screening and computer-aided design have helped identify targets, biomarkers, lead candidates and reduced the time required for advancing an idea from bench to clinic, but it still takes 10-12 years and costs approximately one billion dollars to bring a drug to market globally. Therefore, it is imperative that the strategies to reduce the risk and increase efficiency are carefully selected. In this presentation I would discuss strategies for selecting potential diseases, targets and provide an example of multi-targeted approach to metabolic disorder.
 Colin Barrow, Ph.D.
Colin Barrow, Ph.D.
Chair in Biotechnology, School of Life & Environmental Sciences, Deakin University, Australia
Nano-biotechnology: Omega-3 Oils and Nanofibres
The health benefits of long-chain omega-3 fatty acids are well established, especially for eicosapentaenoic acid (EPA) and docosapentaenoic acid (DHA) from fish and microbial sources. In fact, a billion dollar market exists for these compounds as nutritional supplements, functional foods and pharmaceuticals. This presentation will describe some aspects of our omega-3 biotechnology research that are at the intersection of Nano-biotechnology and oil chemistry. These include the use of lipases for the concentration of omega-3 fats, through immobilization of these lipases on nanoparticles, and the microencapsulation and stabilization of omega-3 oils for functional foods. I will also describe some of our work on the enzymatic production of resolvins using lipoxygenases, and the fermentation of omega-3 oils from marine micro-organisms. Finally, I will describe some of our work on the formation of amyloid fibrils and graphene for various applications in nano-biotechnology.
 Sanjeeva Srivastava, Ph.D.
Sanjeeva Srivastava, Ph.D.
Assistant Professor, Proteomics Lab, IIT-Bombay, India
Identification of Potential Early Diagnostic Biomarkers for Gliomas and Various Infectious Diseases using Proteomic Technologies
The spectacular advancements achieved in the field of proteomics research during the last decade have propelled the growth of proteomics for clinical research. Recently, comprehensive proteomic analyses of different biological samples such as serum or plasma, tissue, CSF, urine, saliva etc. have attracted considerable attention for the identification of protein biomarkers as early detection surrogates for diseases (Ray et al., 2011). Biomarkers are biomolecules that can be used for early disease detection, differentiation between closely related diseases with similar clinical manifestations as well as aid in scrutinizing disease progression. Our research group is performing in-depth analysis of alteration in human proteome in different types of brain tumors and various pathogenic infections to obtain mechanistic insight about the disease pathogenesis and host immune responses, and identification of surrogate protein markers for these fatal human diseases.
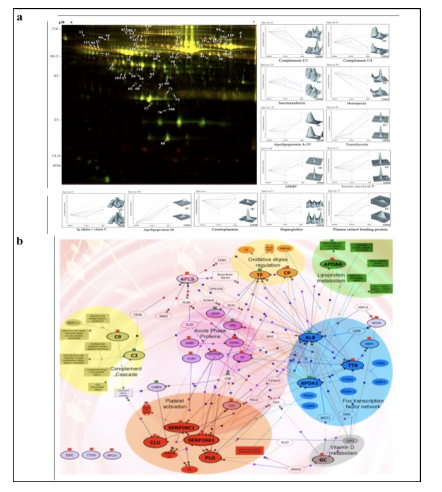
Applying 2D-DIGE in combination with MALDI-TOF/TOF MS we have analyzed the serum and tissue proteome profiles of glioblastoma multiforme; the most common and lethal adult malignant brain tumor (Gollapalli et al., 2012) (Figure 1). Results obtained were validated by employing different immunoassay-based approaches. In serum proteomic analysis we have identified some interesting proteins like haptoglobin, ceruloplasmin, vitamin-D binding protein etc. Moreover, proteomic analysis of different grades (grade-I to IV) of gliomas and normal brain tissue was performed and differential expressions of quite a few proteins such as SIRT2, GFAP, SOD, CDC42 have been identified, which have significant correlation with the tumor growth. While proteomic analysis of cerebrospinal fluid from low grade (grade I & II) vs. high grade (grade III & IV) gliomas revealed modulation of CSF levels of apolipoprotein E, dickkopf related protein 3, vitamin D binding protein and albumin in high grade gliomas. The prospective candidates identified in our studies provide a mechanistic insight of glioma pathogenesis and identification of potential biomarkers. We are also studying the role of JAK/STAT interactome and therapeutic potential of STAT3 inhibitors in gliomas using proteomics approach. Several candidates of the JAK/STAT interactome were identified with altered expression and a significant correlation was observed between STAT3 and PDK1 transcript expression level.
We have also investigated the changes in human serum proteome in different infectious diseases including falciparum and vivax malaria (Ray et al., 2012a; Ray et al., 2012b), dengue (Ray et al., 2012c) and leptospirosis (Srivastava et al., 2012). Although, quite a few serum proteins were found to be commonly altered in different infectious diseases and might be a consequence of inflammation mediated acute phase response signaling, uniquely modulated candidates were identified in each pathogenic infection indicating the some inimitable responses. Further, a panel of identified proteins consists of six candidates; serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I was used to build statistical sample class prediction models employing PLSDA and other classification methods to predict the clinical phenotypic classes and 91.37% overall prediction accuracy was achieved (Figure 2). ROC curve analysis was carried out to evaluate the individual performance of classifier proteins. The excellent discrimination among the different disease groups on the basis of differentially expressed proteins demonstrates the potential diagnostic implications of this analytical approach.
Keywords: Diagnostic biomarkers, Gliomas, Infectious Diseases, Proteomics, Serum proteome
Acknowledgments: This disease biomarker discovery research was supported by Department of Biotechnology, India grant (No. BT/PR14359/MED/30/916/2010), Board of Research in Nuclear Sciences (BRNS) DAE young scientist award (2009/20/37/4/BRNS) and a startup grant 09IRCC007 from the IIT Bombay. The active support from Advanced Center for Treatment Research and Education in Cancer (ACTREC), Tata Memorial Hospital (TMH), and Seth GS Medical College and KEM Hospital Mumbai, India in clinical sample collection process is gratefully acknowledged.
References :
- Ray S, Reddy PJ, Jain R, Gollapalli K. Moiyadi A, Srivastava S. Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139-61, 2011.
- Gollapalli K, Ray S, Srivastava R, Renu D, Singh P, Dhali S, Dikshit JB, Srikanth R, Moiyadi A, Srivastava S. Investigation of serum proteome alterations in human glioblastoma multiforme. Proteomics 12(14): 2378-90, 2012.
- Ray S, Renu D, Srivastava R, Gollapalli K, Taur S, Jhaveri T, Dhali S, Chennareddy S, Potla A, Dikshit JB, Srikanth R, Gogtay N, Thatte U, Patankar S, Srivastava S. Proteomic investigation of falciparum and vivax malaria for identification of surrogate protein markers. PLoS One 7(8): e41751, 2012a.
- Ray S, Kamath KS, Srivastava R, Raghu D, Gollapalli K, Jain R, Gupta SV, Ray S, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Patankar S, Srivastava S. Serum proteome analysis of vivax malaria: An insight into the disease pathogenesis and host immune response. J Proteomics 75(10): 3063-80, 2012b.
- Srivastava R, Ray S, Vaibhav V, Gollapalli K, Jhaveri T, Taur S, Dhali S, Gogtay N, Thatte U, Srikanth R, Srivastava S. Serum profiling of leptospirosis patients to investigate proteomic alterations. J Proteomics 76: 56-68, 2012.
- Ray S, Srivastava R, Tripathi K, Vaibhav V, Srivastava S. Serum proteome changes in dengue virus-infected patients from a dengue-endemic area of India: towards new molecular targets? OMICS 16(10): 527-36, 2012c.
* Correspondence: Dr. Sanjeeva Srivastava, Department of Biosciences and Bioengineering, IIT Bombay, Mumbai 400 076, India: E-mail: sanjeeva@iitb.ac.in; Phone: +91-22-2576-7779, Fax: +91-22-2572-3480
![Figure 2 (a) Western blot analysis of haptoglobin (HP), serum amyloid A (SAA), and clusterin (CLU) from serum samples of healthy control (HC) [n = 12], falciparum malaria (FM) [n = 12], vivax malaria (VM) [n = 12], Leptospirosis (Lep) [n = 6], dengue fever [DF] [n = 6] and non infectious disease control (NIDC:GBM) [n = 12]. Representative blots of the target proteins are depicted along with their respective relative abundance volumes (volume X 104). All the data are represented as mean ± SE. (b) Discrimination of malaria from dengue, leptospirosis and GBM using PLS-DA analysis. PLS-DA scores Plot for FM (blue spheres, n = 8), VM (green spheres, n = 8), DF (red spheres, n = 6), Lep (grey spheres, n = 6) and GBM (brown spheres, n = 8) samples based on 6 differentially expressed proteins (serum amyloid A, hemopexin, apolipoprotein E, haptoglobin, retinol-binding protein and apolipoprotein A-I) identified using DIGE. The axes of the plot indicate PLSDA latent variables t0-t2.](http://www.amritabioquest.org/conference/2013/wp-content/uploads/sites/2/2015/02/sanjeeva-2.jpg)
 Nader Pourmand, Ph.D.
Nader Pourmand, Ph.D.
Director, UCSC Genome Technology Center,University of California, Santa Cruz
Biosensor and Single Cell Manipulation using Nanopipettes
Approaching sub-cellular biological problems from an engineering perspective begs for the incorporation of electronic readouts. With their high sensitivity and low invasiveness, nanotechnology-based tools hold great promise for biochemical sensing and single-cell manipulation. During my talk I will discuss the incorporation of electrical measurements into nanopipette technology and present results showing the rapid and reversible response of these subcellular sensors to different analytes such as antigens, ions and carbohydrates. In addition, I will present the development of a single-cell manipulation platform that uses a nanopipette in a scanning ion-conductive microscopy technique. We use this newly developed technology to position the nanopipette with nanoscale precision, and to inject and/or aspirate a minute amount of material to and from individual cells or organelle without comprising cell viability. Furthermore, if time permits, I will show our strategy for a new, single-cell DNA/ RNA sequencing technology that will potentially use nanopipette technology to analyze the minute amount of aspirated cellular material.
 Kal Ramnarayan, Ph.D.
Kal Ramnarayan, Ph.D.
Co-founder President & Chief Scientific Officer, Sapient Discovery, San Diego, CA, USA
A cost-effective approach to Protein Structure-guided Drug Discovery: Aided by Bioinformatics, Chemoinformatics and computational chemistry
With the mapping of the human genome completed almost a decade ago, efforts are still underway to understand the gene products (i.e., proteins) in the human biological and disease pathways. Deciphering such information is very important for the discovery and development of small molecule drugs as well as protein therapeutics for various human diseases for which no cure exists. As an example, with more than 500 members, the kinase family of protein targets continues to be an important and attractive class for drug discovery. While how many of the members in this family are actually druggable is still to be established, there are several ongoing efforts on this class of proteins across a broad spectrum of disease categories. Even though in general the protein structural topology might looks similar, there are issues with respect selectivity of identified small molecule inhibitors when, the lead molecule discovery is carried out at the ATP binding site. As an added complexity, allosteric modulators are needed for some of the members, but the actual site for such modulation on the protein target can not resolved with uncertainty. In this presentation we will describe a bioinformatics and computational based platform for small molecule discovery for protein targets that are involved in protein-protein interactions as well as targets like kinases and phosphatases. We will describe a computational approach in which we have used an informatics based platform with several hundred kinases to sort through in silico and identify inhibitors that are likely to be highly selective in the lead generation phase. We will discuss the implication of this approach on the drug discovery of the kinase and phosphatase classes in general and independent of the disease category.
 Shigeki Miyamoto, Ph.D.
Shigeki Miyamoto, Ph.D.
Professor, McArdle Laboratory for Cancer Research – UW Carbone Cancer Center
Department of Oncology, School of Medicine and Public Health
University of Wisconsin-Madison
“Inside-out” NF-κB signaling in cancer and other pathologies
The NF-κB/Rel family of transcription factors contributes to critical cellular processes, including immune, inflammatory and cell survival responses. As such, NF-κB is implicated in immunity-related diseases, as well as multiple types of human malignancies. Indeed, genetic alterations in the NF-κB signaling pathway are frequently observed in multiple human malignancies. NF-κB is normally kept inactive in the cytoplasm by inhibitor proteins. Extracellular ligands can induce the release of NF-κB from the inhibitors to allow its migration into the nucleus to regulate a variety of target genes. NF-κB activation is also induced in response to multiple stress conditions, including those induced by DNA-damaging anticancer agents. Although precise mechanisms are still unclear, research from our group has revealed a unique nuclear-to-cytoplasmic signaling pathway. In collaboration with bioengineers, clinicians and pharmaceutical industry, our lab has developed new methods to analyze primary cancer patient samples and identified several compounds with different mechanisms that mitigate this cell survival pathway. Further contributions from other labs have also revealed additional mechanisms and molecular players in this “inside-out” signaling pathway and expanded its role in other physiological and pathological processes, including B cell development, premature aging and therapy resistance of certain cancers. Our own new findings, along with these recent developments in the field, will be highlighted.

John Stanley, Satheesh Babu, Ramacahandran T and Bipin Nair
Pt-Pd decorated TiO2 nanotube array for the non-enzymatic determination of glucose in neutral medium
Rapidly expanding diabetic population and the complications associated with elevated glycemic levels necessitates the need for a highly sensitive, selective and stable blood glucose measurement strategy. The high sensitivity and selectivity of enzymatic sensors together with viable manufacturing technologies such as screen-printing have made a great social and economic impact. However, the intrinsic nature of the enzymes leads to lack of stability and consequently reduces shelf life and imposes the need for stringent storage conditions. As a result much effort has been directed towards the development of ‘enzyme-free’ glucose sensors (Park et al. 2006). In this paper, a non-enzymatic amperometric sensor for selective and sensitive direct electrooxidation of glucose in neutral medium was fabricated based on Platinum-Palladium (Pt–Pd) nanoparticle decorated titanium dioxide (TiO2) nanotube arrays. Highly ordered TiO2 nanotube arrays were obtained using a single step anodization process (Grimes C A and Mor G K 2009) over which Pt–Pd nanoparticles where electrochemically deposited. Scanning Electron Microscopy (SEM) analysis revealed the diameter of the TiO2 nanotubes to be approximately 40 nm. Elemental analysis after electrochemical deposition confirms the presence of Pt–Pd. Electrochemical characterization of the sensor was carried out using cyclic voltammetry technique (−1.0 to +1.0V) in phosphate buffer saline (PBS) pH 7.4. All further glucose oxidation studies were performed in PBS (pH 7.4). The sensor exhibited good linear response towards glucose for a concentration range of 1 μM to 20mM with a linear regression coefficient of R = 0.998. The electrodes are found to be selective in the presence of other commonly interfering molecules such as ascorbic acid, uric acid, dopamine and acetamidophenol. Thus a nonenzymatic sensor with good selectivity and sensitivity towards glucose in neutral medium has been developed.
The global healthcare scene of which the pharmaceutical industry and its products are integral components is today at the cross roads. The high and unaffordable costs of drug research with estimates of over 1 billion dollars for every new drug discovered and developed, the very low success rates, the high degree of obsolescence due to undesirable adverse drug reactions, the decline in the development pipeline of new drugs, patent expiries leading to generic competition and the public’s disillusionment with use of chemicals for human consumption as drugs have all significantly contributed to the problems of this lifeline industry. The strategy adopted by the large R&D based Corporations to get bigger and bigger through mergers and acquisitions to improve cost-effectiveness and productivity of R&D has so far not worked effectively. Consequently, one of the recent trends in healthcare, articulated by many experts is to look for alternate or even complementary approaches to reduce the impact of rising costs of drugs on healthcare. Various new strategies for drug discovery such as the use of Natural Products especially medicinal plants are being actively pursued by healthcare planners and providers. Side by side, traditional systems of medicine whether from the oriental countries or the western nations are also having a serious relook to understand their usefulness in healthcare. To achieve its legitimate position in the healthcare scenario, it is essential to scientifically validate their claimed utility through appropriate and systematic research efforts including pre-clinical and clinical studies. In addition to their own use as medicines, knowledge on the Indian Traditional Medicines can be used as a platform for new drug discovery. The huge potential for carrying out systematic R&D programs for new Drug Discovery based on natural products and possible strategies to realise them in the coming decades will be explained in this presentation.
 Ramani A. Aiyer, Ph.D., MBA
Ramani A. Aiyer, Ph.D., MBA
Principal, Shasta BioVentures, San Jose, CA, USA
New Drug R&D in India: Challenges & Opportunities
New drug discovery and development has become a global endeavor, with Western big pharmaceutical companies farming out more and more chemistry and biology research to Asia, particularly India and China. During the last decade, several Indian pharmaceutical companies have embarked on ambitious R&D programs, with slow but steady progress in developing new chemical / molecular entities. The Indian government has also made a strong commitment to promote innovation and entrepreneurship in the biotechnology sector. The first part of the talk will focus on a case study showing the entire process of discovery and development of a new drug recently launched for Rheumatoid Arthritis. We will then address the challenges of conducting innovative R&D in India and actions necessary to overcome them. The second part of the talk will make the case for developing Ayurvedic drug formulations for the Western / Global markets, again using the example of Rheumatoid Arthritis (Aamavaata). Ayurveda takes a holistic approach to disease diagnosis and therapy based on interactions among body type (prakriti), tri-doshas (three body humors), sapta-dhatus (seven tissues) and malas (excretions). The drugs prescribed are usually herbo-mineral formulations comprising multiple medicinal plants and / or metals. The manufacturing processes date back to Ayurvedic texts several thousand years old, and are compiled in the Ayurvedic Pharmacopeia. Also, the treatment modalities and drug formulations are “personalized” to fit different patient types, based on the holistic diagnoses mentioned earlier. There is a tremendous need to establish a sound basis for Ayurvedic drug discovery R&D for the modern world. We must find a scientific and ethical way to leverage the vast body of anecdotal and possibly retrospective data on patients undergoing Ayurvedic treatment. Combined with in vitro and in vivo biological data on Ayurvedic herbo-mineral formulations, the adoption of stringent manufacturing practices, and designing sound clinical trials to establish the safety and efficacy, India has a golden opportunity to expand the reach of Ayurvedic drugs into Western / Global medical practice.







